Quality Management System
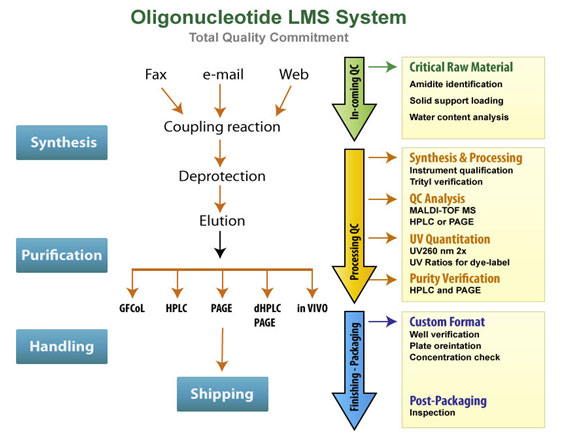
Producing high quality oligonucleotides requires a rigorous quality control system in place. All the reagents used in our oligo synthesis process are sourced from reliable suppliers and checked for quality prior to use. During synthesis, we apply trityl monitoring of oligonucleotides to control the coupling efficiency at each cycle. All oligonucleotides are routinely analyzed by analytical HPLC or PAGE for purity and characterized by MALDI-TOF, LC-MS or ESI mass spectrometry. Finally, oligos are quantitated twice by UV spectrophotometry to provide an accurate measure of yield.
Customer Support
With nearly 30 years of experience in oligonucleotide synthesis, we have the expertise to synthesize the most technically challenging custom biomolecules. Our Molecular Biologists and Chemists are prepared to assist you with experimental design, application support, and troubleshooting.
Guaranteed Quality
As part of our long-standing commitment to quality, we employ the following standards for quality assurance:
- All nucleic acid synthesis products (including RNA, siRNAs, DNA) are analyzed by mass spectrometry and quantified at least twice by uv spectrophotometry (unless no post-synthesis processing is requested)
- MALDI-TOFMS: for all standard DNA, RNA and siRNA synthesis
- HPLC, PAGE, RNase Free HPLC and PAGE: for purification of DNA and RNA products
- UPLC or CGE: for quality analysis when appropriate
- QC methods and documentation are available upon request
QC documentation is available upon request including:
- Mass Spectrometry traces
- UPLC or CGE analysis
- Duplex or individual strand purity analysis
- Certificate of Analysis
- Endotoxin certification
- Custom QC documentation
The following applies to mid-scale synthesis projects (200 mg - 10 g):
- HPLC purification is highly recommended to achieve optimal purity
- Tangential Flow Filtration (TFF) is utilized for premium salt removal
- Batch records are maintained for all synthesis projects
- Endotoxin testing is included on all mid-scale orders
- MS, UPLC, endotoxin results, and a certificate of analysis are supplied