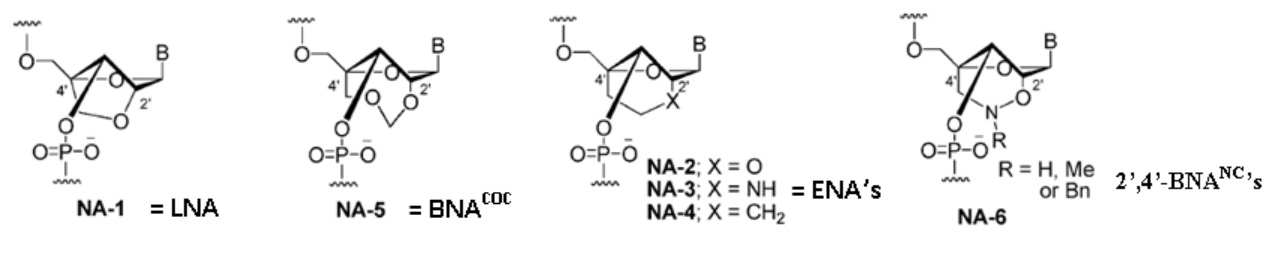
2’,4’-BNA (LNA) with a five-membered bridged structure is insufficiently resistant to nucleases, and fully modified 2’,4’-BNA (LNA) oligonucleotides do not have the flexibility required for efficient triplex formation.
BNA analogues with increased steric bulk and less conformational restriction were developed. 2’,4’-BNACOC has a seven membered bridged structure and exhibits dramatically improved nuclease resistance.
NA-5 = BNACOC: This modification conversely affects duplex stability [i.e., duplexes formed with this nucleic acid analogue are less stable than those formed by 2’,4’-BNA (LNA)].
ENA, with a six-membered bridged structure, has slightly lower duplex-forming ability and significantly higher nuclease resistance than 2’,4’-BNA (LNA).
Triplex formation with ENA provided variable results compared to that of 2’,4’-BNA (LNA).
2’,4’-BNANC, has a six-membered bridged structure with a unique structural feature (N-O bond) in the sugar moiety. The bridged moiety was designed to have a nitrogen atom, which can:
(i) Act as a conjugation site
(ii) (I(ii) mprove duplex and triplex stability by lowering repulsions between the negatively charged backbone phosphates
The nitrogen atom on the bridge can be functionalized by hydrophobic and hydrophilic groups, by steric bulk, or by appropriate functional moieties to:
(i) Control affinity toward complementary strands
(ii) Regulate resistance against nuclease degradation, and
(iii) Allows synthesis of functional molecules designed for specific applications in genomics.
In addition to the above possibilities, the N-O bond could be cleaved under appropriate reductive conditions to modulate the hybridizing properties of 2’,4’-BNANC.
I. Duplex formation with RNA based on Tm
|
| Target ssRNA: |
5’-r(AGCAAAAAACGC)-3’ |
|
| Duplex: |
5’-r(AGCAAAAAACGC)-3’
3’-d(TCGTTTTTTGCG)-5’ |
Duplex formation with 2’,4’-BNANC[NH] chimer:
|
| |
2’,4’-BNANC[NH] |
Tm |
ΔTm |
ΔTm/BNA |
| d Oligo |
GCGTTTTTTGCT |
45 |
|
|
| 1 BNA |
GCGTTTTTTGCT |
51 |
+6 |
+6 |
| 3 BNAs |
GCGTTTTTTGCT |
64 |
+19 |
+6.3 |
| 3 BNAs |
GCGTTTTTTGCT |
61 |
+16 |
+5.3 |
| 6 BNAs |
GCGTTTTTTGCT |
83 |
+38 |
+6.3 |
Duplex formation with 2’,4’-BNANC[NH] chimer:
| |
2’,4’-BNANC[NH] |
Tm |
ΔTm |
ΔTm/BNA |
| d Oligo |
GCGTTTTTTGCT |
45 |
|
|
| 1 BNA |
GCGTTTTTTGCT |
50 |
+5 |
+5 |
| 3 BNAs |
GCGTTTTTTGCT |
63 |
+18 |
+6 |
| 3 BNAs |
GCGTTTTTTGCT |
59 |
+14 |
+4.7 |
| 6 BNAs |
GCGTTTTTTGCT |
80 |
+35 |
+5.8 |
Duplex formation with 2’,4’-BNA (LNA) chimer:
| |
2’,4’-BNANC[NH] |
Tm |
ΔTm |
ΔTm/BNA |
| d Oligo |
GCGTTTTTTGCT |
45 |
|
|
| 1 BNA |
GCGTTTTTTGCT |
52 |
+7 |
+7 |
| 3 BNAs |
GCGTTTTTTGCT |
62 |
+17 |
+5.7 |
| 3 BNAs |
GCGTTTTTTGCT |
60 |
+15 |
+5 |
| 6 BNAs |
GCGTTTTTTGCT |
80 |
+35 |
+5.8 |
| Note 1: |
A dOligo with one LNA monomer binds the strongest. |
| Note 2: |
A dOligo with multiple 2’,4’-BNANC[NMe] monomers bind with higher affinities to RNA. |
| Note 3: |
A dOligo with multiple 2’,4’-BNANC[NMe] monomers bind with similar affinities to RNA when compared to LNA. |
Conclusion: |
2’,4’-BNANC[NH] is more specific to RNA. LNA and 2’,4’-BNANC[NMe] are similar.
|
II. Duplex formation with DNA based on Tm
|
| Target ssDNA: |
5’-d(AGCAAAAAACGC)-3’ |
|
| Duplex: |
5’-d(AGCAAAAAACGC)-3’
3’-d(TCGTTTTTTGCG)-5’ |
Duplex formation with 2’,4’-BNANC[NH] chimer:
|
| |
2’,4’-BNANC[NH] |
Tm |
ΔTm |
ΔTm/BNA |
| d Oligo |
GCGTTTTTTGCT |
50 |
|
|
| 1 BNA |
GCGTTTTTTGCT |
51 |
+1 |
+1 |
| 3 BNAs |
GCGTTTTTTGCT |
55 |
+18 |
+1.7 |
| 3 BNAs |
GCGTTTTTTGCT |
57 |
+14 |
+2.3 |
| 6 BNAs |
GCGTTTTTTGCT |
73 |
+23 |
+3.8 |
Duplex formation with 2’,4’-BNANC[NMe] chimer:
| |
2’,4’-BNANC[NH] |
Tm |
ΔTm |
ΔTm/BNA |
| d Oligo |
GCGTTTTTTGCT |
50 |
|
|
| 1 BNA |
GCGTTTTTTGCT |
49 |
-1 |
-1 |
| 3 BNAs |
GCGTTTTTTTGCT |
51 |
+1 |
+0.3 |
| 3 BNAs |
GCGTTTTTTGCT |
50 |
0 |
0 |
| 6 BNAs |
GCGTTTTTTGGCT |
61 |
+11 |
+1.8 |
Duplex formation with 2’,4’-BNA(LNA) chimer:
| |
2’,4’-BNANC[NH] |
Tm |
ΔTm |
ΔTm/BNA |
| d Oligo |
GCGTTTTTTGCT |
50 |
|
|
| 1 BNA |
GCGTTTTTTGCT |
53 |
+3 |
+3 |
| 3 BNAs |
GCGTTTTTTTGCT |
56 |
+6 |
+2 |
| 3 BNAs |
GCGTTTTTTGCT |
54 |
+4 |
1.3 |
| 6 BNAs |
GCGTTTTTTGGCT |
67 |
+17 |
+2.8 |
Note 1: |
A dOligo with one LNA monomer binds the strongest. |
| Note 2: |
dOligos with multiple 2’,4’-BNANC[NH] monomers bind with higher affinities to DNA than dOligos with 2’,4’-BNANC[NMe]. |
| Note 3: |
A dOligo with multiple 2’,4’-BNANC[NMe] monomers bind with higher affinities to DNA when compared to LNA oligos. |
Conclusion: |
Oligos with several 2’,4’-BNANC[NMe] bind more strongly to DNA when compared to oligos with 2’,4’-BNANC[NMe] or LNA monomers.
|
Tm Values of Duplexes
|
| Formed by 2’,4’-BNANC- and 2’,4’-BNA (LNA)-Modified Oligonucleotides with Complementary ssRNA Containing a Single-Mismatch Basea,b |
| Tm (ΔTm= Tm (mismatch) -Tm (match)(oC) |
| T (oligonucleotide) |
X ) A (match) |
U |
G |
C |
| GCGTTTTTTGCT |
45 |
33(-12) |
43(-3) |
30(-15) |
| 2',4'-BNANC[NH] GCGTTTTTTGCT |
51 |
37 (-14) |
46(-5) |
34 (-17) |
| 2',4'-BNANC[NMe] GCGTTTTTTGCT |
50 |
37 (-13) |
48 (-2) |
34 (-16) |
| 2',4'-BNA (LNA) GCGTTTTTTGCT |
52 |
39 (-13) |
47 (-5) |
35 (-17) |
a Modified oligonucleotides, 5'-d(GCGTTTTTTGCT)-3'; Target ssRNA strand, 3'-(CGCAAXAAACGA)-5'. b Conditions: 10 mM sodium phosphate buffer (pH 7.2) containing 100 mM NaCl; strand concentration) 4 µM.
Conclusion:
ΔTm values in the Table show that any mismatched base in the target RNA strand resulted in a substantial decrease in the Tm of duplexes formed with 2’,4’-BNANC-modified oligonucleotides.
ΔTm values of duplexes formed with 2’,4’-BNANC[NH]-modified oligonucleotide having T-U, T-G, and T-C arrangements are -14, -5, -17 °C.
These are lower than those of the corresponding natural DNA-RNA duplexes (ΔTm= -12, -3, and -15 °C, respectively).
Similar to those exhibited by duplexes formed with the 2’,4’-BNA (LNA) oligonucleotide (ΔTm= -13, -5, and -17 °C, respectively).
The mismatch discrimination profiles of 2’,4’-BNANC[NMe] and [NBn] were also similar to that of 2’,4’-BNA (LNA), except in the case of the T-G arrangement for the 2’,4’-BNANC[NMe] derivative, as shown in the Table.
Thus, it appears that 2’,4’-BNANC not only exhibits high-affinity RNA selective hybridization, but is also highly selective in recognizing bases.
III. Triplex Formation and Triplex Stability.
|
|
Applications in antigene and gene repair technologies require the formation of stable triplexes at physiological pH. Rahman et al in 2007 showed that 2’,4’-BNANC[NH] has triplex-forming characteristics when compared to those of 2’,4’-BNA (LNA) and ENA. Details of triplex formation by three 2’,4’-BNANC analogues (2’,4’-BNANC[NH], [NMe], and [NBn]) in the absence and presence of divalent metal (Mg2+) are described here.
Triplex-forming ability of the 2’,4’-BNANC analogues against a 21 bp double-stranded DNA (dsDNA) in the absence of divalent metal is summarized in next Table.
A single modification of the triplex-forming oligonucleotide (TFO) with 2’,4’-BNANC[NH] (TFO) increased the Tm of the triplex by 11 °C, which is equal to that exhibited by the triplex of 2’,4’-BNA (LNA)-modified TFO.
The Tm of the triplex formed by the corresponding 2’,4’-BNANC-[NMe]-TFO is 5 °C higher than that of natural-TFO, and that of 2’,4’-BNANC[NBn]-TFO is equal to that of the natural oligonucleotide. In the case of 2’,4’-BNANC[NH], increasing the number of modifications (TFOs) greatly enhanced triplex thermal stability (ΔTm/mod. = +6.2 to +9 °C), which is similar to or even higher than that of the corresponding 2’,4’-BNA (LNA)-TFOs (ΔTm/mod. = +4.9 to +8.7 °C).
The thermal stability of the triplex formed by the corresponding 2’,4’-BNANC[NMe]-TFOs, although significantly higher than that obtained with natural DNA-TFO, is lower than that of 2’,4’-BNANC[NH]- and 2’,4’-BNA (LNA)-TFOs.
Modification with 2’,4’-BNANC-[NBn] units did not affect the thermal stability of the triplex; that is, the thermal stability was similar to that of natural DNA-TFO. The lower stability of the 2’,4’-BNANC-[NMe] and [NBn] triplexes might be due to unfavorable steric interaction with dsDNA, induced by the methyl or benzyl groups.
(Rahman, S. M. A.; Seki, S.; Obika, S.; Haitani, S.; Miyashita, K.; Imanishi, T. Angew. Chem., Int. Ed. 2007, 46, 4306{c}).
|
| a Target dsDNA: |
5’-d(GCTAAAAAGAAAGAGAGATCG)-3’
3’-d(CGATTTTTCTTTCTCTCTAGC)-5’ |
|
underlined portion indicates the target site for triplex formation. b Conditions: 7 mM sodium phosphate buffer (pH 7.0) containing 140 mM KCl; strand concentration = 1.5 µM. c Data from Rahman et al. 2007.
Conclusion:
Oligonucleotides with 2’,4’-BNANC[NH] monomers form more stable TFOs when compared to 2’,4’-BNANC[NMe] and LNAs.
Proper spacing of the monomers appears to make the binding stronger.
The best placement of BNANC[NH] monomers for triplex probes is:
|
|
|
|
This design appears to be the best for the design of probes that form stable triplexes under physiological conditions!
This is similar or close to LNA oligos. 2’,4’-BNANC[NH] oligos form more stable TFOs.
|
Molecular models of Triplexes
|
 |
 |
|
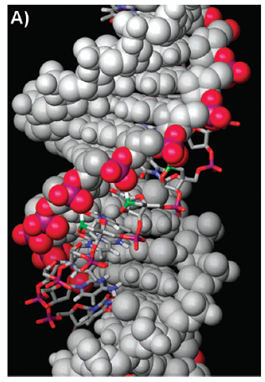
Molecular modeling of a parallel-motif triplex formed by 2’,4’-BNANC[NH]-TFO with dsDNA.
BNA: d(TTTTTmCTTTmCTmCTmCT). (Source: Rahman et al., 2008).
(A) The overall view of the triplex is depicted.
The dsDNA is shown as a gray CPK model with the phosphate backbone of the purine strand colored red and purple.
TFO is shown as a colored tube model with three 20,40-BNANC nitrogens represented in green.
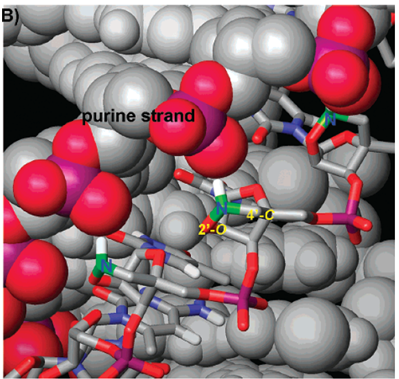
(B) Expanded view of the area of the triplex containing three consecutive 20,40-BNANC[NH] residues.
Note that the 20,40-BNANC nitrogens are very close to the phosphate moiety of the purine strand indicating ionic interactions of the compounds.
|