Can antisense oligonucleotides containing bridged nucleic acids (BNAs) be used for COVID-19 therapies?
Recent research suggests that bridged nucleic acid (BNA) antisense oligonucleotides (ASOs) enable the design of SARS-CoV-2 (COVID-19) targeting therapeutics. Many vaccines and drugs are now worldwide in development to stop COVID-19 infections and to treat infected people. In general, drug design is mostly based on protein targets since 3D structures for virus proteins are more readily available.
The SARS-CoV-2 (COVID-19) virus requires the frameshift stimulation element (FSE) for a balanced expression of essential viral proteins.
The coronavirus, SARS-CoV-2, has a positive sense (+), single-stranded RNA (ssRNA) genome. The first two open reading frames (ORFs) 1a and 1b encode for non-structural proteins. The ORFs include coding for the RNA-dependent polymerase and partial overlap. The precise stoichiometric expression of ORF 1a and ORF 1b throughout the viral replication cycles is vital for viral fitness. An essential regulatory mechanism regulates the two reading frames expression ratio called a -1 programmed ribosomal frameshifting (- PRF). A structured RNA motif at the 3’-end of ORF 1a called the frameshift stimulation element (FSE) initiates the PRF mechanism. FSE directs elongating ribosomes to randomly shift their reading frames by one base in the 5’-direction. The frameshift enables the readthrough past the ORF 1a stop codon needed to maintain appropriate levels of ORF 1a to ORF 1ab expression. FSE has a 5’-heptanucleotide “slippery site” UUUAAAC followed by an RNA element thought to form a three-stem pseudoknot.
Zhang et al. recently reported the 3D structure of the synthetic 28 kDA FSE from the SARS-CoV-2 RNA genome. The research team used cryo-electron microscopy and model-building to determine the 3D structure of the SARS-CoV-2 FSE with a 6.9 Å (0.69 nm) resolution.
Structural model of the 28-kDa Frameshift Stimulation Element from the SARS-CoV-2 RNA Genome
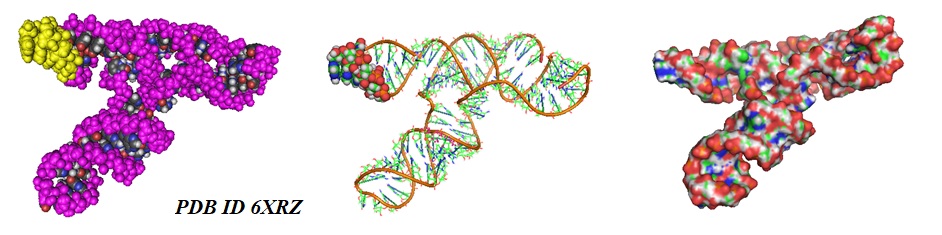
Figure 1: Models of the Frameshift Stimulation Element from the SARS-CoV-2 RNA Genome (6XRZ). A space fill model (left), a cartoon model (middle), and a surface model (right) is shown. The sequence TTAAAC is highlighted in yellow and as space fill residues.
GTTTTTAAACGGGTTTGCGGTGTAAGTGCAGCCCGTCTTACACCGTGCGGCACAGGCACTAGTACTGATGTCGTATACAGGGCTTTTG
SARS-CoV-2 FSE targeting BNA ASOs
The team utilized bridged nucleic acid (BNA/LNA) modified antisense oligonucleotides for the targeted disruption of the RNA frames shifting element. A few selected BNA ASOs were also synthesized with phosphorothioate (PS) internucleotide linkages. Binding pockets identified in the FSE 3D structure as targets to interrupt the element allowed the guided design of ASOs, including BNA ASOs.
Frameshift Stimulation Element Sequences and targeting ASOs
>pdb|6XRZ|A Chain A, Frameshift Stimulation Element from the SARS-CoV-2 RNA Genome
GTTTTTAAACGGGTTTGCGGTGTAAGTGCAGCCCGTCTTACACCGTGCGGCACAGGCACTAGTACTGATGTCGTATACAGGGCTTTTG
MW390853.1
GTTTTTAAACGGGTTCGCGGTGTAAGTGCAGCCCGTCTTACACCGTGCGGCACAGGCACTAGTACTGATGTCGTATACAGGGCTTTTG
LC528233.2
GTTTTTAAACGGGTTTGCGGTGTAAGTGCAGCCCGTCTTACACCGTGCGGCACAGGCACTAGTACTGATGTCGTATACAGGGCTTTTG
----AATTTGCCCAAGCGCCA----------------------------------------------TACAGCATATGTCCC(S2D)-
----AATTTGCCCAAACGCCA(SS2)-------------------------------TGATCATGACTACA(S3D1)-----------
----------------------------------------------------GTCCGTGATCATGACT(S3D2)--------------
NNN = BNAs
The UUUAAAC sequence is highly conserved in coronavirus genomes.
ASO and BNA ASOs used in the study.
|
ASOs
|
Sequence
|
Linkage
|
|
Stem 2 Disruptor (S2D)
|
5′-+CCCTG+TA+TA+CGACA+T-3′
|
-
|
|
Stem 3 Disruptor-1 (S3D1)
|
5′-+A+C+ATCAGTACT+A+G+T-3′
|
PS linkages
|
|
Stem 3 Disruptor-2 (S3D2)
|
5′-+T+C+AGTACTAGTG+C+C+T+G-3′
|
PS linkages
|
|
Slippery site 2 (SS2)
|
5′-+A+C+C+GCGAACCCGTT+T+A+A-3′
|
PS linkages
|
Reference
Ref: Zhang K, Zheludev IN, Hagey RJ, Wu MT, Haslecker R, Hou YJ, Kretsch R, Pintilie GD, Rangan R, Kladwang W, Li S, Pham EA, Bernardin-Souibgui C, Baric RS, Sheahan TP, D Souza V, Glenn JS, Chiu W, Das R. Cryo-electron Microscopy and Exploratory Antisense Targeting of the 28-kDa Frameshift Stimulation Element from the SARS-CoV-2 RNA Genome. bioRxiv [Preprint]. 2020 Jul 20:2020.07.18.209270. doi: 10.1101/2020.07.18.209270. PMID: 32743589; PMCID: PMC7386510. [PMC]
---...---