PCR based methods as well as in-situ hybridization multiplexing methods when used for pathogen detection allow the simultaneous analysis of multiple nucleic acid sequences in a single reaction. Multiplexing can potentially save a lot of time, cost and labor but requires precise work and often involves many complicated steps. The design of general primers enables the development of universal methods for the detection of multiple species during a single analysis run.
Real-time PCR (quantitative PCR, qPCR) is now a very well-established detection, quantification, and typing method for different microbial agents in a variety of diagnostic settings. However, to make PCR successful, there are specific issues in qPCR that developers and users of this technology must address. Kralik and Ricchi in 2017 published an essential guide that describes the use of the correct terminology and definitions, the understanding of the principle of PCR, difficulties with interpretation and the presentation of data, as well as the limitations of qPCR in different areas of microbial diagnostics and parameters necessary for qPCR performance.
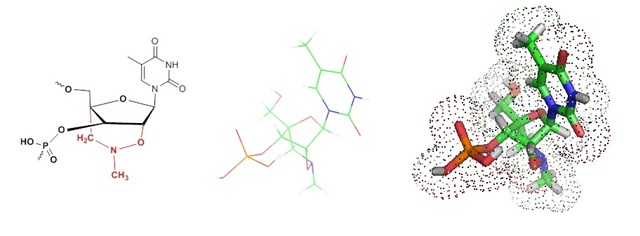
Oligonucleotides modified with bridged nucleic acid (BNAs) residues have stronger affinities for their complementary targets than natural nucleic acid bases and are known to avoid dimer formation and mismatch hybridization. The use of BNAs achieving enhanced and efficient priming.
Barghouthi SA in 2011 reported the development of a universal method allowing the detection of 101 tested isolates by producing one or more PCR products from each isolate. Sequencing of strains and aligning the resulting sequences using BLAST enabled the identification of the detected bacterial species.
To enable detection of microbes in the gut of animals and possibly humans, Guillen et al. in 2016 showed that 16S primer sets enabled the analysis of bacteria in the colon of rats. The use of selected universal primers provided knowledge of the abundance of microorganisms and the bacterial diversity in rat colon biopsies.
Sun et al. in 2008 showed that oligonucleotide pentamer primer pairs modified with BNAs enabled the amplification of genomic DNA demonstrating that the use of bridged nucleic acid pentamers as universal primer pairs together with suspension array genotyping allowed the identification of multiple distinct genes or species with a single amplification procedure. Sun et al. thereby demonstrated that BNA based modified pentamer-based PCR are very useful for pathogen detection and identification. Furthermore, Levin et al. in 2006 were able to show that the incorporation of BNAs at the 5’-prime end of PCR primers increased the Phred Q30 score during DNA sequencing dramatically.
Biosynthesis Inc. offers BNA modified oligonucleotides.
Reference
Barghouthi SA. A universal method for the identification of bacteria based on general PCR primers. Indian J Microbiol. 2011;51(4):430-44. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3209952/
Guillen IA, Camacho H, Tuero AD, et al. PCR Conditions for 16S Primers for Analysis of Microbes in the Colon of Rats. J Biomol Tech. 2016;27(3):105-12. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4920503/
Kralik P, Ricchi M. A Basic Guide to Real Time PCR in Microbial Diagnostics: Definitions, Parameters, and Everything. Front Microbiol. 2017;8:108. Published 2017 Feb 2. doi:10.3389/fmicb.2017.00108. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5288344/
Joshua D. Levin, Dean Fiala, Meinrado F. Samala, Jason D. Kahn, Raymond J. Peterson; Position-dependent effects of locked nucleic acid (LNA) on DNA sequencing and PCR primers, Nucleic Acids Research, Volume 34, Issue 20, 1 November 2006, Pages e142, https://doi.org/10.1093/nar/gkl756
Lopez I, Ruiz-Larrea F, Cocolin L, et al. Design and evaluation of PCR primers for analysis of bacterial populations in wine by denaturing gradient gel electrophoresis. Appl Environ Microbiol. 2003;69(11):6801-7. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC262258/
Sun Z, Chen Z, Hou X, Li S, Zhu H, Qian J, et al. (2008) Locked Nucleic Acid Pentamers as Universal PCR Primers for Genomic DNA Amplification. PLoS ONE 3(11): e3701. https://doi.org/10.1371/journal.pone.0003701. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0003701
---…---