Minor Groove Binders or MGBs are crescent-shaped molecules that selectively bind non-covalently to the minor groove of DNA, a shallow furrow in the DNA helix. Binding to DNA with specific sequences usually takes place by a combination of directed hydrogen bonding to base pair edges.
Natural Minor Groove Binder Molecules
Duocarmycin A and its analogs are naturally occurring antitumor agents. These molecules belong to a group of DNA minor groove binding molecules that exhibit AT-sequence selective adenine-N3-alkylations. Duocarmycins have been isolated from Streptomyces bacteria, and a number of these compounds have already been used during clinical trials to test them for their potential as anticancer drugs.
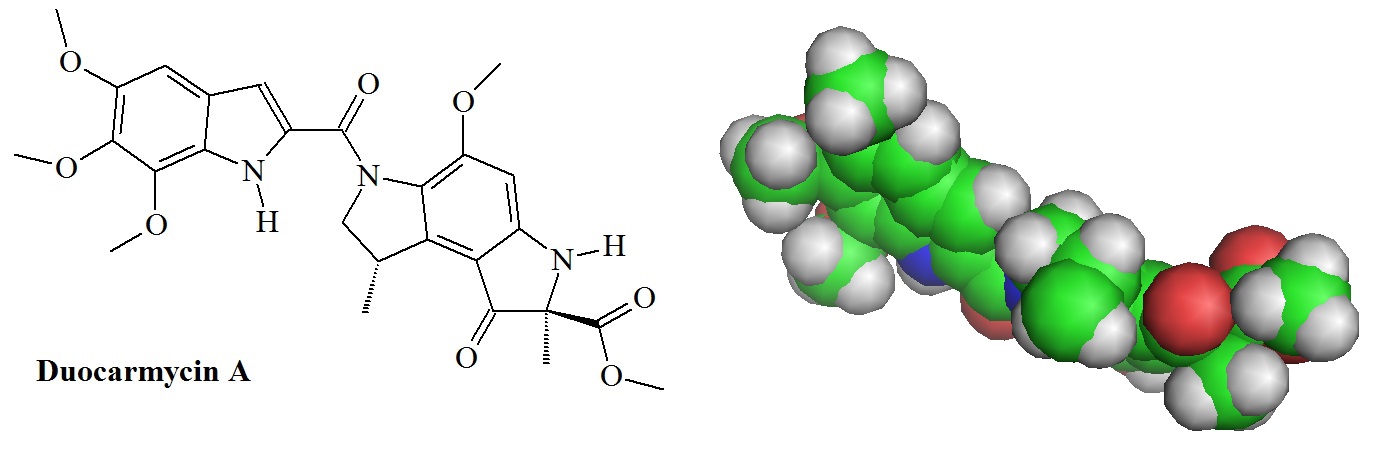
Figure 1: Structure and molecular model of duocarmycin A (Source: PDB 107D). Duocarmycins are known to derive their potent antitumor activity through sequence-selective minor groove alkylation of N3 adenine in double-stranded DNA. Duocarmycins belong to a new class of antitumor antibiotics containing a unique spiro-cyclo-propyl-hexadienone moiety responsible for DNA alkylation. The water-soluble derivative, KW-2189, has broad-spectrum antitumor activity in a series of experimental tumor models and is presently in Phase II clinical trials.
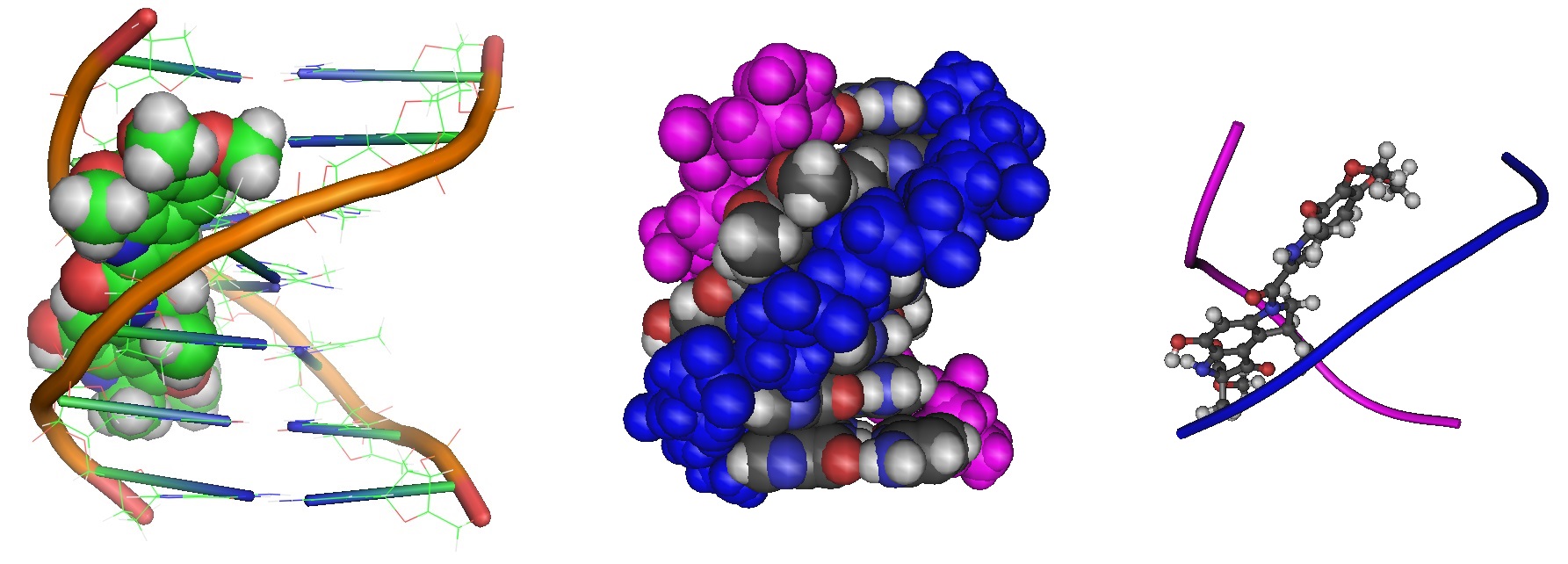
Figure 2: Different renderings of molecular models of a duocarmycin A-DNA duplex complex (Source: PDB 107D). The model is based on the solved solution structure using a combined NMR-molecular dynamics study including NOE based intensity refinement by Lin and Patel in 1995. The structure revealed that the antitumor antibiotic duocarmycin A binds covalently to the minor groove N-3 position of adenine with sequence specificity for the 3'-adenine in a d(A-A-A-A) tract in duplex DNA. The adenine ring is protonated during duocarmycin adduct formation resulting in charge delocalization over the purine ring system. The model shows a minimally perturbed right-handed duplex in the B-form helix and duocarmycin A is positioned within the walls of the minor groove.
Hoechst dyes are Minor Groove Binders
Hoechst dyes are a family of blue fluorescent dyes useful for the staining of DNA. These bis-benzimide dyes were developed by the Hoechst AG. The three dyes Hoechst 33342, 333580, and 333258 are the most commonly used.
The bisbenzimide Hoechst 33258 and its derivatives are starting molecules for the design of minor groove binders. These molecules are widely used for staining DNA in cells and belong to a class of small molecules that preferably bind to AT-rich sequences. The dye Hoechst 33258 is known to induce apoptosis and enhance over expression of transgenes.
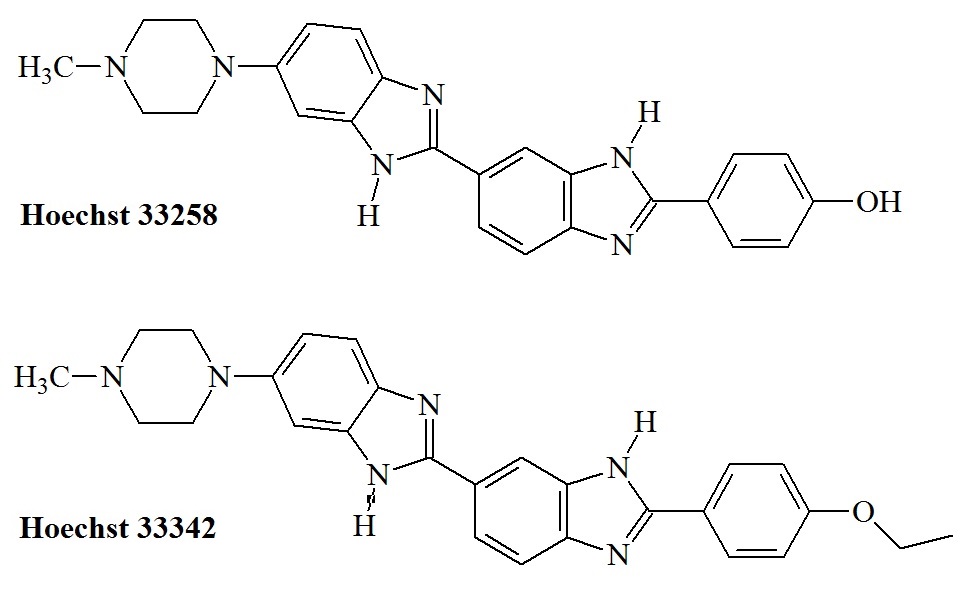
Figure 3: Chemical structures of the dyes Hoechst 33258 and 33342.
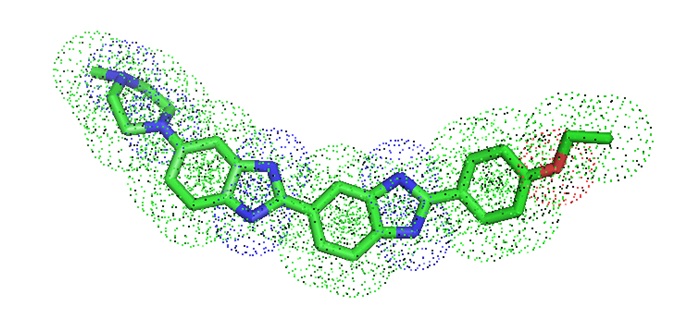
Figure 4: Molecular model of the dye Hoechst 33342.
When cells are exposed to alkylating agents DNA lesions are the result. Because mammalian cells have DNA repair mechanisms, most of these lesions are repaired. But some of these lesions can persist and can now induce point mutations subsequently leading to carcinogenic conversions. One such persistent lesion, O6-Ethylguanine (e6G), is caused by the exposure of DNA to N-ethyl-N-nitrosourea.
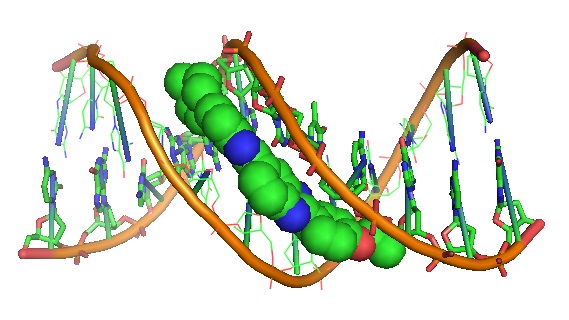
Figure 5: Molecular structure of d(CGC[e6G]AATTCGCG in complex with Hoechst 33342 containing O6-ethyl-G-C base pairs (Source: PDB 129D). O6-ethyl-G (e6G) is an important DNA lesion caused by the exposure of cells to alkylating agents such as N-ethyl-N-nitrosourea.
Three dye-duplex DNA structures were solved at approximately 2.0 A resolution. The structures showed that the Hoechst molecule lies squarely at the central AATT site with the ends approaching the G4-C21 and the G16-C9 base pairs. However, the two independent e6G-C base pairs in the DNA duplex were found to adopt different base pairing types. The e6G4-C21 base pair has a configuration similar to a normal Watson-Crick base pair, except with bifurcated hydrogen bonds between e6G4 and C21, and the ethyl group is in the proximal orientation. The e6G16-C9 base pair is reported to adopt a wobble configuration and the ethyl group is in the distal orientation.
The alkylation products of guanine are shown in figure 6.
Figure 6: Structures of 6-O-methylguanine, 6-O-ethylguanine, and guanine.
The dealkylating enzyme 6-O-methylguanine methyltransferase can remove methyl groups from the O6 position of guanine.
O6-Ethylguanine (e6G) is a relatively persistent alkylation lesion caused by the exposure of DNA to N-ethyl-N-nitrosourea. Exposure to N-nitrosamides leads to the formation of N- and O-alkylation products in cellular DNA, for example to N-ethyl-N-nitrosourea or N-methyl-N-nitrosourea. O6-ethylguanine can cause G:C to A:T transition mutation. This has been observed in codon 12 (GGA) of the H-ras gene in rat mammary and skin tumors induced by N-methyl-N-nitrosourea. O6–Alkylated Guanine in DNA can be repaired by the repair protein O6-alkylguanine-DNA alkyltransferase. As the size of the alkyl residue increases the repair can also occur via the nucleotide excision repair pathway. The enzyme catalyzes the transfer of the alkyl group to Cys 145 at the active site of the protein and the formation of S-alkylcysteine irreversibly inactivates the protein.
Reference
Akira Asai, Keiichi Yano, Tamio Mizukami and Hirofumi Nakano; Characterization of a Duocarmycin-DNA Adduct-recognizing Protein in Cancer Cells. Cancer Res November 1 1999 (59) (21) 5417-5420.
https://www.sciencedirect.com/topics/neuroscience/hoechst-stain
https://en.wikipedia.org/wiki/Hoechst_stain
http://www.biotech.illinois.edu/sites/biotech.illinois.edu/files/uploads/Hoechst.pdf
Lin CH, Patel DJ.; Solution structure of the covalent duocarmycin A-DNA duplex complex. J Mol Biol. 1995 Apr 21;248(1):162-79. https://www.ncbi.nlm.nih.gov/pubmed/7731041.
---...---