Terminal deoxynucleotidyl transferase (TdT; EC 2.7.7.31) is a DNA polymerase that elongates DNA strands template-independently and incorporates both ribo- and deoxy-ribonucleotides and many other unnatural nucleoside triphosphates into oligonucleotides in vitro. TdT performs DNA synthesis using only single-stranded DNA as the nucleic acid substrate. TdT was discovered in 1960 and purified from a calf thymus gland. F.J Bollum showed that a calf thymus polymerase preparation catalyzes primer-dependent reactions in the presence of homogeneous polydeoxynucleotides.
In animals with a backbone (vertebrates), the highly conserved TdT adds diversity to the immune repertoire by adding nucleotides to the V(D)J recombination junction sites of immunoglobulin and T-cell receptor genes. The added nucleotides are known as N regions. The enzyme incorporates nucleotides that increase antigen receptor diversity randomly. The ~1014 different immunoglobulins and ~1018 unique T cell antigen receptors generated this way are required for antigen neutralization. G. Jäger, in 1981, developed a method for detecting terminal transferase in individual cells. The approach uses an unlabeled antibody based enzyme method to classify acute lymphoblastic leukemias that have variably differentiated T-cells.
TdT is part of the polymerase family called pol X, a subclass of an ancient nucleotidyltransferase (NT) superfamily. Other nucleic acid polymerases such as DNA polymerase β (pol β), DNA polymerase λ and DNA polymerase, CCA-adding enzymes, and poly(A) polymerases also belong to this NT superfamily. The expression of TdT appears to be restricted to thymic T cells and a small Ig negative cell fraction in the bone marrow. These findings are useful for the classification of lymphoid tumors. The following figures show molecular models for terminal deoxynucleotidyl transferase.
Recently the enzyme has been harnessed for the production of digital DNA. Lee at al. designed a de novo enzymatic synthesis strategy for data storage which uses the template-independent polymerase terminal deoxynucleotidyl transferase in kinetically controlled conditions. In this approach information is stored in transitions between non-identical nucleotides of DNA strands. For the production of strands representing user-defined content, nucleotide substrates are added iteratively to generate short homopolymeric extensions whose lengths are controlled by apyrase-mediated substrate degradation.
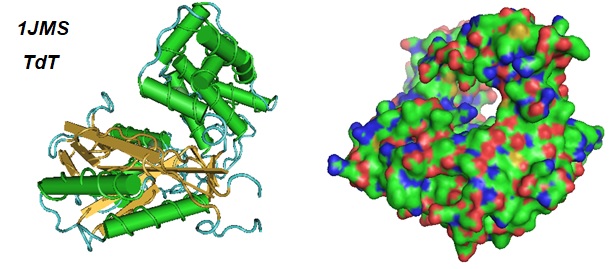
Figure 1: Models of the catalytic core of murine terminal deoxynucleotidyl-transferase (TdT) resulting from the crystal structure at 2.35 A resolution (PDB ID 1JMS; Delarue et al.). The secondary structure model (Left) and the surface model (Right) is depicted. The protein contains a typical DNA polymerase beta-like fold locked in a closed form. Solving of the structure showed that the substrates and two divalent ions in the catalytic site are positioned in TdT in a manner like the human DNA polymerase beta ternary complex. These observations suggested a common two metal ions mechanism of nucleotidyl transfer in these proteins as proposed by Steitz and Steitz in 1993. The inability of TdT to accommodate a template strand can be explained by steric hindrance at the catalytic site caused by a long lariat-like loop, absent in DNA polymerase beta.

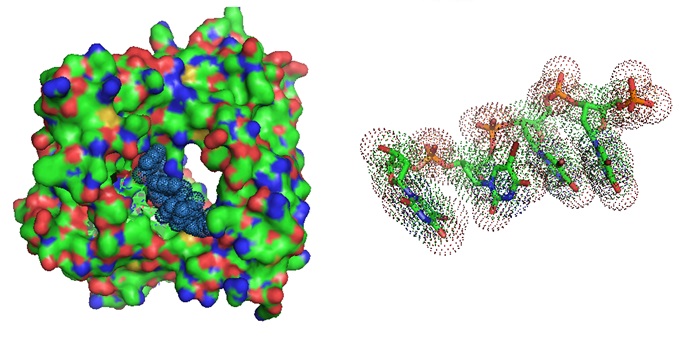
Figure 2: Different models of the Terminal Deoxynucleotidyltransferase Short Isoform and its substrate [PDB ID 1KDH]. The binary complex of murine terminal deoxynucleotidyl transferase with a primer single stranded DNA is illustrated. TRANSFERASE-DNA COMPLEX; Mol_id: 1; Molecule: 5'-D(P*(BRU)P*(BRU)P*(BRU)P*(BRU))-3'; Chain: D; Mol_id: 2; Molecule: Terminal deoxynucleotidyltransferase short isoform; Chain: A; EC number: 2.7.7.31. [PubMed]
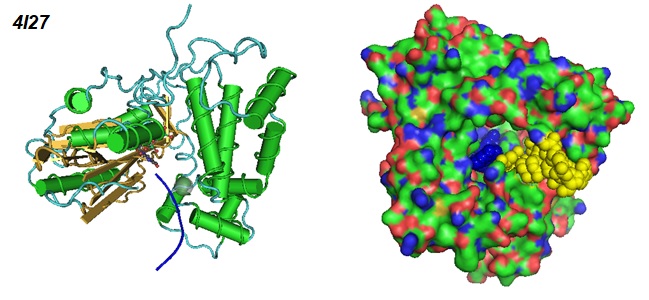
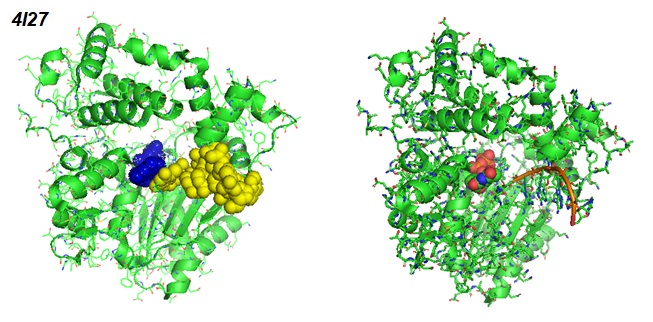
Figure 3: Different views of the Ternary Complex Of Mouse Tdt With SsDNA And Incoming Nucleotide (Gouge et al. 2013. PDB ID 4I27).
Reference
Bollum FJ. Oligodeoxyribonucleotide primers for calf thymus polymerase. J Biol Chem. 1960 May; 235:PC18-20. PMID: 13802336.
Bollum, F. J. (1960) Calf thymus polymerase.J. Biol. Chem. 235, 2399–2403. [PubMed]
Bollum, F. J.; Terminal Deoxynucleotidyl Transferase. The Enzymes, Volume 10, 1974, Pages 145-171. [Sciencedirect]
Delarue M, Boulé JB, Lescar J, Expert-Bezançon N, Jourdan N, Sukumar N, Rougeon F, Papanicolaou C. Crystal structures of a template-independent DNA polymerase: murine terminal deoxynucleotidyltransferase. EMBO J. 2002 Feb 1;21(3):427-39. [PMC]
Deng, G.R., Wu, R., Terminal transferase: Use in the tailing of DNA and for in vitro mutagenesis, Meth. Enzymol., 100, 96-116, 1983. [Pubmed]
Eschenfeldt, W.H., et al.; Homopolymeric tailing. Meth. Enzymol., 152, 337-342, 1987. [Pubmed] [ScienceDirect]
Frohman, M.A., et al., Rapid Production of Full-Length cDNAs from Rare Transcripts: Amplification Using a Single Gene-Specific Oligonucleotide Primer. Proc. Natl. Acad. Sci. USA, 85, 8998-9002, 1988. [PMC]
Gaastra, W., Klemm, P., Radiolabeling of DNA with 3' terminal transferase, Methods in Molecular Biology, 1985, 2, 269-271. [PubMed]
Gorczyca, W., et al., Detection of DNA strand breaks in individual apoptotic cells by the in situ terminal deoxynucleotidyl transferase and nick translation assays, Cancer Res., 53, 1945-1951,1993. [Cancer Research]
Gouge J, Rosario S, Romain F, Beguin P, Delarue M. Structures of intermediates along the catalytic cycle of terminal deoxynucleotidyltransferase: dynamical aspects of the two-metal ion mechanism. J Mol Biol. 2013 Nov 15;425(22):4334-52. [PMC]
Heiss M, Kellner S. Detection of nucleic acid modifications by chemical reagents. RNA Biol. 2017 Sep 2;14(9):1166-1174. [PMC]
Igloi, G.L., Schiefermayr, E.; Enzymatic addition of fluorescein-or biotin-riboUTP to oligonucleotides results in primers suitable for DNA sequencing and PCR, BioTechniques, 15, 486-497, 1993. [Pubmed]
Kumar, A., et al., Nonradioactive labeling of synthetic oligonucleotide probes with terminal deoxynucleotidyl transferase, Anal. Biochem., 169, 376-382, 1988. [Pubmed]
Lee, H.H., Kalhor, R., Goela, N. et al. Terminator-free template-independent enzymatic DNA synthesis for digital information storage. Nat Commun 10, 2383 (2019). [Nature]
Motea EA, Berdis AJ. Terminal deoxynucleotidyl transferase: the story of a misguided DNA polymerase. Biochim Biophys Acta. 2010 May;1804(5):1151-1166. [PMC]
Steitz TA, Steitz JA. A general two-metal-ion mechanism for catalytic RNA. Proc Natl Acad Sci U S A. 1993 Jul 15;90(14):6498-6502. [PMC]
Terminal deoxynucleotidyl transferase (TdT) wiki
Tu, C.-P.D., Cohen, S.N.; 3’-end labeling of DNA with [α-32P]-cordycepin-5’-triphosphate, Gene, 10, 177-183, 1980. [Pubmed]
Vincent, C., Tchen, P., Cohen-Solal, M., & Kourilsky, P.; Synthesis of 8-(2,4-dinitrophenyl-2,6-aminohexyl)aminoadenosine-5'-triphosphate: Biological properties and potential uses, Nucleic Acids Res., 10, 6787-6796, 1982. (rATP-DNP) [PMC]
---...---