Capping and decapping of mRNA are cellular events during gene expression. The chemical nature of the 5′ end of RNA appears to determine the stability of RNA during RNA processing, and localization, as well as the translation efficiency. Therefore it has been proposed that this type of modification provides an additional layer of gene regulation called “epitranscriptomic” gene regulation.
Enzymes that are known to decap mRNA belong to two different protein families, the Nudix pyrophosphohydrolases and the HIT family of pyrophosphatases.
The nucleoside diphosphates linked to moiety-X (NUDIX) hydrolases belong to a superfamily of enzymes conserved throughout all species. Originally this protein family was called the MutT protein family. The MutT protein family hydrolyzes a nucleoside diphosphate linked to some other moiety, X. Therefore researchers proposed to call this protein family the “nudix” family instead of the “MutT protein family.”
Nudix decapping enzymes hydrolyze the phosphodiester bond between the beta and the alpha phosphates in a metal-dependent manner resulting in a 5’-monophosphate RNA (pRNA) and a 7-methyl-guanosine diphosphate (m7-GDP).
However, the HIT decapping enzyme family hydrolyzes the phosphodiester bond between the gamma and beta phosphates. The result is a 5′-diphosphate-RNA (ppRNA) and a 7-methyl-guanosine monophosphate (m7GMP).
In general, the HIT protein family based hydrolyzes is independent of divalent metal ions. The so-called scavenger decapping enzyme DcpS is a member of the HIT family of pyrophosphatases. The DcpS scavenger decapping enzyme clears the cell of residual cap structures that would otherwise accumulate. This reaction is the final step of the 3’ to 5’ end mRNA decay pathway. Therefore it is suggested that DcpS influences the pool of available cap-binding proteins and impacts their downstream functions. Dcps appears to be a regulatory protein. However, more research is needed to clarify this.
Recent findings indicate that bacterial as well as eukaryotic transcripts are capped with cellular cofactors. Specific RNA polymerases add the cofactors during transcription initiation. Mitochondrial RNA binding proteins RNAP) also cap transcripts with ADP-containing cofactors. However, the role of universal RNAP-catalyzed capping is not yet clear.
Nudix hydrolases occur in all organism and hydrolyze a wide range of organic pyrophosphates. Nudiz enzymes hydrolyze nucleoside di- and triphosphates, dinucleoside and diphosphoinositol polyphosphates, nucleotide sugars and RNA caps with varying degrees of substrate specificity. Some of these enzymes degrade potentially mutagenic, oxidized nucleotides. Other enzymes of these protein family control the levels of metabolic intermediates and signaling compounds.
In 2017, Bird et al. showed that NAD+, NADH, and dpCoA function as initiating nucleotides (NCINs) for de novo transcription initiation. The research group reported the crystal structures of transcription initiation complexes containing NCIN-capped RNAs and the mechanism and structural basis of NCIN capping, suggesting that NCIN-mediated “ab initio capping” may occur in all organisms, also reported as the capping of cellular RNAs.
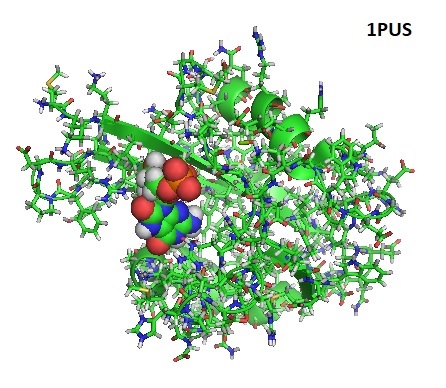
Figure 1: Solution Structure of the MutT Pyrophosphohydrolase Complexed with Mg(2+) and 8-oxo-dGMP. PDB 1PUS.
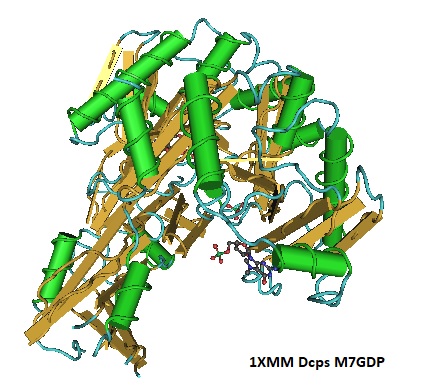
Figure 2: Structure Of Human Dcps Bound To M7GDP. PDB 1XMM.
Metabolically labile nucleoside phosphoramidates can be incorporated into nucleotide analogs either chemically or enzymatically. These types of nucleotide analogs have found applications as prodrugs and are called “ProTides”. ProTides can be cleaved by the human histidine triad nucleotide binding protein 1 (hHint1) to expose the nucleotide monophosphate which evate the highly selective nucleoside kinases that limit the use of nucleosides as prodrugs.
A combination of enzyme kinetic methods with X-ray crystallography and isothermal titration calorimetry allows exploration of the energetics of substrate binding and studying the structural basis for catalytic efficiency of ProTides.
Reference
Abeygunawardana C., Weber D. J., Gittis A., Frick D. N., Lin J., Miller A.-F., Bessman M. J., Mildvan A. S.; Solution structure of the MutT enzyme, a nucleoside triphosphate pyrophosphohydrolase. (1995) Biochemistry 34, 14997–15005.
Bird JG, Zhang Y, Tian Y, Panova N, Barvík I, Greene L, Liu M, Buckley B, Krásný L, Lee JK, Kaplan CD, Ebright RH, Nickels BE. The mechanism of RNA 5? capping with NAD+, NADH and desphospho-CoA. Nature. 2016 Jul 21;535(7612):444-7. doi: 10.1038/nature18622. PubMed PMID: 27383794; [PubMed]
Maize KM, Shah R, Strom A, Kumarapperuma S, Zhou A, Wagner CR, and Finzel BC; A Crystal Structure Based Guide to the Design of Human Histidine Triad Nucleotide Binding Protein 1 (hHint1) Activated ProTides. Mol Pharm. 2017 Nov 6;14(11):3987-3997. doi: 10.1021/acs.molpharmaceut.7b00664. Epub 2017 Oct 26.
Massiah MA, Saraswat V, Azurmendi HF, Mildvan AS; Solution structure and NH exchange studies of the MutT pyrophosphohydrolase complexed with Mg(2+) and 8-oxo-dGMP, a tightly bound product. Biochemistry (2003) 42 p.10140-54
Maurice J. Bessman, David N. Frick and Suzanne F. O'Handley; The MutT Proteins or “Nudix” Hydrolases, a Family of Versatile, Widely Distributed, “Housecleaning” Enzymes. The Journal of Biological Chemistry 271, 25059-25062.
McLennan AG; The Nudix hydrolase superfamily. Cell Mol Life Sci. 2006 Jan;63(2):123-43. [Pubmed]
Julius C, Riaz-Bradley A, Yuzenkova Y. RNA capping by mitochondrial and multi-subunit RNA polymerases. Transcription. 9(5):292-297. doi: 10.1080/21541264.2018.1456258. Epub 2018 Apr 25. PubMed PMID: 29624107; PubMed Central PMCID: PMC6150613. [Pubmed]
Carreras-Puigvert J, Zitnik M, Jemth AS, Carter M, Unterlass JE, Hallström B, Loseva O, Karem Z, Calderón-Montaño JM, Lindskog C, Edqvist PH, Matuszewski DJ, Ait Blal H, Berntsson RPA, Häggblad M, Martens U, Studham M, Lundgren B, Wählby C, Sonnhammer ELL, Lundberg E, Stenmark P, Zupan B, Helleday T. A comprehensive structural, biochemical and biological profiling of the human NUDIX hydrolase family. Nat Commun. 2017 Nov 16;8(1):1541. doi: 10.1038/s41467-017-01642-w. PubMed PMID: 29142246; [PubMed]
Wulf MG, Buswell J, Chan SH, Dai N, Marks K, Martin ER, Tzertzinis G, Whipple JM, Corrêa IR Jr, Schildkraut I. The yeast scavenger decapping enzyme DcpS and its application for in vitro RNA recapping. Sci Rep. 2019 Jun 13;9(1):8594. doi: 10.1038/s41598-019-45083-5. PubMed PMID: 31197197; [PubMed]
---...---