Neuroblastoma represents the most common cancer for infants less than 1 year old and ~90% are diagnosed at younger than 5 years of age. In humans, nervous system is comprised of ‘central nervous system’ (brain, spinal cord) and ‘peripheral nervous system’ (ganglia, nerves located externally to spinal cord or brain). The peripheral system is composed of ‘somatic nervous system’ (cranial nerves originating from brainstem that controls the head and neck anatomical structures) and ‘autonomic nervous system’ (ganglionic neurons connected to brain that involuntarily controls physiological functions by heart, bladder, etc). The autonomic nervous system is divided into ‘sympathetic nervous system’ (controls “fight or flight” response) and ‘parasympathetic nervous system’ (controls salivation, digestion, heart rate, etc). Neuroblastoma occurs in the nerve tissues of sympathetic nervous system (i.e. paraspinal ganglia, adrenal medulla) (Maris, 2010). Consequently, the lesions can appear as a mass in various regions (ex. neck, abdomen, chest, etc.) with wide-ranging symptoms, making the diagnosis of neuroblastoma difficult.
For neuroblastoma, the overall 5-year survival rate remains <40% (Eposito et al, 2017). In another report, a moderate increase in the survival rates (52% for 1975-77 to to 74% for 1999-2005) was documented—primarily attributed to the increased cure rate for the benign type (Maris, 2010). Nearly half of all diagnosed are classified as ‘high risk’ group for recurrence. Depending on the type of risk group to which a neuroblastoma patient is assigned, different treatment may follow. For the ‘low risk’ group with localized tumor, surgery is recommended, whereas for the ‘high risk’ group involving metastasis to bone (or bone marrow), treatments may include surgery, dose-intensive chemotherapy with hematopoietic stem cell transplantation, radiotherapy and immunotherapy. As such, the ability to diagnose accurately is critical as it directly impacts risk stratification as well as therapy.
In trying to classify patients into different risk groups (low, high, intermediate, ultra-high), multiple prognostic factors are considered, which include the age of patient, clinical stage, tumor differentiation, and tumor histology. To help identify the stage, ‘Omics’ data (i.e. proteomics, genomics, metabolomics) are increasingly being utilized. The ability to sequence the entire genome has provided insight on additional layer of information, ex. gene copy number variation, amplification, deletion, genetic variant (ex. single nucleotide polymorphism). Through high-throughput Omics analysis, other relevant information such as mRNA/protein expression level, post-translational modification and metabolites can be acquired.
Neuroblastoma is thought to be a disease of developing tissues that arise from the precursor cells (Hoehner et al, 1996). Genetic analysis has shown that hereditary neuroblastoma, which exhibits autosomal dominant mode of inheritance, is associated with the activating mutation in ALK (ana-plastic lymphoma kinase) oncogene (Mossé et al., 2008) and inactivation of homeobox gene PHOX2B (Trochet et al., 2004). Genome wide association study (GWAS) has implicated several other genes (ex. FLJ22536, BARD1). In ~20% of the cases, N-myc gene is amplified, which is associated with advanced stage neuroblastoma (Brodeur et al., 1984; Lee et al., 1984). N-myc belongs to a family of human proto-oncogenes related to v-myc—i.e. c-myc, N-myc, l-myc. V-myc is an oncogene encoded by avian myelocytomatosis virus (MC29), which causes neoplasm in chickens (myelocytomas, tumors of kidney and liver). In patients with Burkitt lymphoma, a chromosomal translocation places c-myc gene under the control of immunoglobulin promoter, resulting in elevated expression.
In human tissues, three different types of circular DNAs have been identified, i.e. ring chromosome, large extrachromosomal circular DNA (ecDNA), and small extrachromosomal circular DNA (eccDNA). The presence of circular DNA in 93 neuroblastoma samples was computationally inferred by applying algorithms to detect circularity in genome sequencing data (Koche et al., 2019). To increase the sensitivity of detection, the investigators at Sloan Kettering Cancer Center first depleted genomic DNA via nuclease treatment before conducting ‘circle sequencing’ (Circle-seq). Circle sequencing utilizes circularized DNA template, which yields tandem repeats of sequences that are then aligned to determine the consensus sequence. With ‘SMRT-seq’ (single molecule real time sequencing) using fluorescent phospho-linked nucleotides, the sequences could be obtained in real time.
In neuroblastoma, no ring chromosomes were found although ecDNA (mean size 680 kbp; ~0.8 copies per tumor) and eccDNA (mean size 2.4 kbp; ~5600 copies per tumor) were detected. Whereas ecDNA may contain the entire genes, eccDNAs generally contains partial genes, with DNA circularization occurring in both coding and noncoding regions. Intriguingly, the chromosomal region encompassing N-myc gene was highly circularized; other genes circularized include proto-oncogene JUN or MDM2 and transcription factor SOX11 or TAL2.
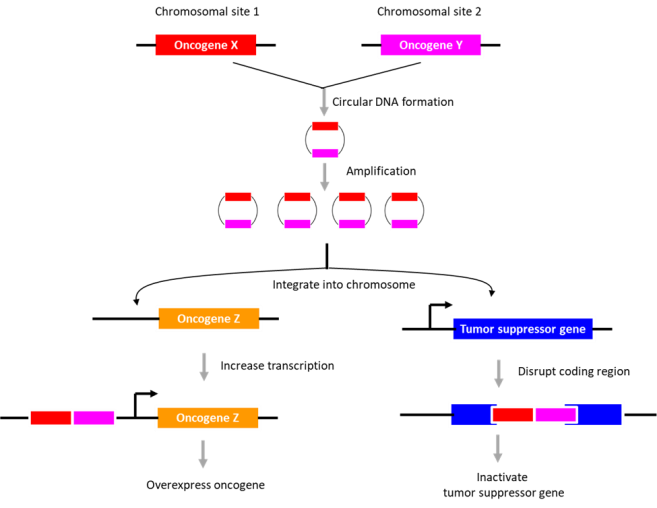
The potential role of DNA circularization in tumorigenesis was investigated. Examination of circle junction sequence revealed microhomologies (minimally 5 bp), indicating that the DNA circles may have arisen during ‘microhomology-mediated DNA repair’. The data suggested that DNA circularization may be necessary but not sufficient for genomic amplification. Also, its role in gene overexpression appeared unlikely. As an alternative, its role in genome remodeling was examined. The extrachromosomal circular DNAs were found to be chimeras consisting of genomic sequences derived from distinct chromosomes (~2.2 chimeric segments for eccDNA; ~4.8 chimeric segments for ecDNA). Further, they described an eccDNA (containing a region in chromosome 2 encompassing the N-myc gene), which is partly integrated into chromosome 13, disrupting DLCK1 gene. Thus, the authors suggest that the extrachromosomal circular DNAs may integrate into various sites in the genome to disrupt tumor suppressor gene function or enhance proto-oncogene expression. The presence of “circle-derived rearrangements” correlated with poor survivability, indicative of its potential use as a diagnostic marker for neuroblastoma.
Bio-Synthesis, Inc. specializes in oligonucleotide modification and provides an extensive array of chemically modified nucleoside analogues (over ~200) including bridged nucleic acid (BNA). A number of options are available to label oligonucleotides (DNA or RNA) with fluorophores either terminally or internally as well as conjugate to peptides. It recently acquired a license from BNA Inc. of Osaka, Japan, for the manufacturing and distribution of BNANC, a third generation of BNA oligonucleotides. To meet the demands of therapeutic application, its oligonucleotide products are approaching GMP grade. Bio-Synthesis, Inc. has recently entered into collaborative agreement with Bind Therapeutics, Inc. to synthesize miR-21 blocker using BNA for triple negative breast cancer. The BNA technology that we offer provides superior, unequalled advantages in base stacking, binding affinity, aqueous solubility and nuclease resistance. It also improves the formation of duplexes and triplexes by reducing the repulsion between the negatively charged phosphates of the oligonucleotide backbone. Its single-mismatch discriminating power was especially useful for diagnosis (ex. FISH using DNA probe). More importantly, BNA oligonucleotide exhibits lesser toxicity than other modified nucleotides for clinical application.
https://www.biosyn.com/oligonucleotide-modification-services.aspx
References
Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM. Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. (1984) Science. 224:1121–4. PMID: 6719137 DOI: 10.1126/science.6719137
Esposito MR, Aveic S, Seydel A, Tonini GP. Neuroblastoma treatment in the post-genomic era. (2017). J Biomed Sci. 24:14. PMID: 28178969 doi: 10.1186/s12929-017-0319-y
Hoehner JC, Gestblom C, Hedborg F, Sandstedt B, Olsen L, Pahlman S. A developmental model of neuroblastoma: differentiating stroma-poor tumors’ progress along an extra-adrenal chromaffin lineage. (1996) Lab Invest. 75:659–75. PMID: 8941212
Koche RP, Rodriguez-Fos E, Helmsauer K, Burkert M, MacArthur IC, Maag J, et al. Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. (2019) Nat Genet. PMID: 31844324 DOI: 10.1038/s41588-019-0547-z
Lee WH, Murphree AL, Benedict WF. Expression and amplification of the N-myc gene in primary retinoblastoma. (1984) Nature 309:458-60. PMID: 6728001 DOI: 10.1038/309458a0
Maris JM. Recent advances in neuroblastoma. (2010) N Engl J Med. 362:2202-11. PMID: 20558371 doi: 10.1056/NEJMra0804577
Mossé YP, Laudenslager M, Longo L, et al. Identification of ALK as a major familial neuroblastoma predisposition gene. Nature. (2008) 455:930–5. PMID: 18724359 PMCID: PMC2672043 DOI: 10.1038/nature07261
Trochet D, Bourdeaut F, Janoueix-Lerosey I, et al. Germ-line mutations of the paired-like homeobox 2B (PHOX2B) gene in neuroblastoma. (2004) Am J Hum Genet. 74:761–4. PMID: 15024693 PMCID: PMC1181953 DOI: 10.1086/383253