What are circular RNAs?
Circular RNAs (circRNAs) are classified as non-coding RNAs (ncRNAs). CircRNAs are covalently closed RNA molecules. Initially considered a splicing error, circRNAs appear to have essential roles in gene regulation. Thousands of circRNAs are expressed from human genomes.
CircRNAs are found in the cytoplasm, are evolutionary conserved, and are relatively stable compared to their linear versions; for example, circRNAs are stable against exonucleolytic decay. The average half-life of endogenously produced 3′-5′-linked circRNA varies from 19 to 24 hours but can be up to 48 hours. However, linear mRNAs have an average lifetime of only 4 to 9 hours. Holdt et al. suggested their use as therapeutic agents and targets.
Generally, circRNAs are generated via "back-splicing" or exon skipping of precursor mRNAs (pre-mRNAs). The 'back-splicing' structure is primarily formed via the junction of a downstream 3′-splice site with an upstream 5′-splice site (head-to-tail splicing). The spliceosome fuses a splice donor site in a downstream exon to a splice acceptor site in an upstream exon.
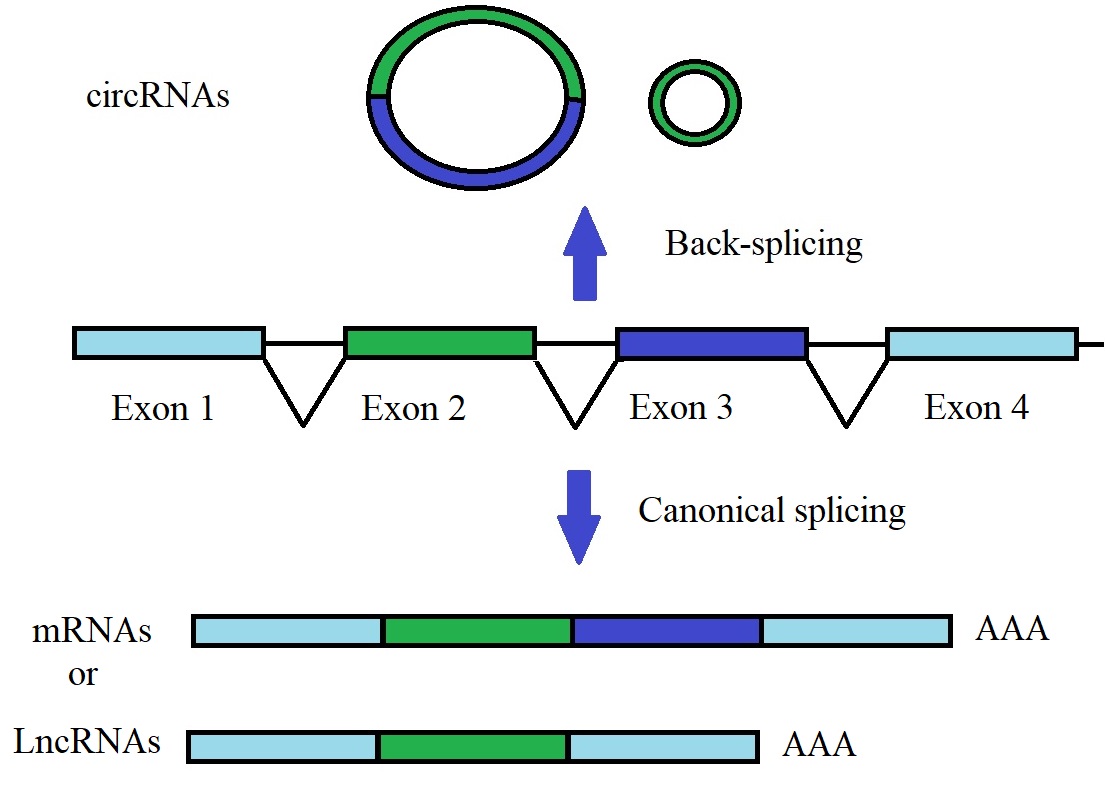
Figure 1: Back-splicing and canonical splicing of a single pre-mRNA.
A single pre-mRNA can be back-spliced with the 5'-terminus upstream of exon 2 ligated to the 3'-terminus of downstream exon 3 generating a circular RNA. During canonical splicing, the exons of the pre-mRNA can be joined colinearly to form mRNAs or lncRNAs (Adapted from Yu & Kuo 2019).
CircRNAs can resist exonucleolytic degradation by RNase R. Also, exon skipping results in a restricted lariat structure that can promote cyclization.
Based on their composition, currently, circRNAs are divided into four categories:
1. Exonic circular RNAs (EcircRNAs; ~80% or more);
2. Circular intronic RNAs (ciRNAs);
3. Exon–intron circRNAs (EIciRNAs); and
4. tRNA intronic circular RNAs (tricRNAs) formed by tRNA introns.
CiRNAs and EIciRNAs are predominantly localized in the nucleus, where they may regulate gene transcription.
Exons from the flanking regions of a gene form so called "read-through" circRNAs via back-splicing. CircRNAs can participate in several physiological and pathological processes.
Biological roles of circular RNA:
(A) RNA binding protein (RBP)-mediated circularization.
(B) Intron pairing-driven circularization.
(C) Lariat-driven circularization.
(D) tRNA intronic circular RNAs (tricRNAs) are formed during the process of pre-tRNA splicing.
(E) Exon-intron circular intronic RNAs (EIciRNAs) can interact with U1 small nuclear ribonucleoproteins. This interaction increases the transcription of their host genes by binding with RNA pol II. ciRNAs and the RNA pol II complex can directly interact and regulate parental gene transcription.
(F) Circularization can compete with canonical splicing.
(G) EcircRNAs can trap mRNAs.
(H) CircRNAs can sponge miRNAs.
(I) CircRNAs can act as protein sponges.
(J) CircRNAs can be translated into peptides or proteins.
Holdt et al. describe two fundamentally different modes of circRNA biogenesis:
(1) Cotranscriptional back-splicing within the linear pre-mRNA and
(2) Posttranscriptional back-splicing from within already excised exon(s)- and intron(s)-containing lariats and intra-lariat back-splicing occurs physically separated from the maturing linear mRNA molecule.
Research studies of circRNA biogenesis are ongoing and bioinformatic algorithms for mapping circRNAs will surely constantly improve. For the over-expression of circRNAs in cells, DNA constructs utilize parts of native introns with inverted repeats or artificially cloned sequences in reverse complementary orientation adjacent to circulation exons.
Reference
Chen, R., Wang, S.K., Belk, J.A. , Amaya, L., Li, Z., Cardenas, A., Abe, B.T., Chen, C.K., Wender, P.A., Cahng, H.Y.; Engineering circular RNA for enhanced protein production. Nat Biotechnol (2022). [PubMed]
Enuka, Y., Lauriola, M., Feldman, M. E., Sas-Chen, A., Ulitsky, I., and Yarden, Y. (2016). Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 44, 1370–1383. [PMC]
Hansen, T. B. (2018). Improved circRNA identification by combining prediction algorithms. Front. Cell Dev. Biol. 6:20. [PMC]
Holdt LM, Kohlmaier A, Teupser D. Circular RNAs as Therapeutic Agents and Targets. Front Physiol. 2018 Oct 9;9:1262. [PMC]
Jeck, W. R., Sorrentino, J. A., Wang, K., Slevin, M. K., Burd, C. E., Liu, J., et al. (2013). Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157. [PMC]
Schwanhausser, B., Busse, D., Li, N., Dittmar, G., Schuchhardt, J., Wolf, J., et al. (2011). Global quantification of mammalian gene expression control. Nature 473, 337–342. [PubMed]
Szabo, L., and Salzman, J. (2016). Detecting circular RNAs: bioinformatic and experimental challenges. Nat. Rev. Genet. 17, 679–692. [PMC]
Yu CY, Kuo HC. The emerging roles and functions of circular RNAs and their generation. J Biomed Sci. 2019 Apr 25;26(1):29. [PMC]
Peng Zhang, Xiao-Ou Zhang, Tingting Jiang, Lingling Cai, Xiao Huang, Qi Liu, Dan Li, Aiping Lu, Yan Liu, Wen Xue, Peng Zhang, Zhiping Weng, Comprehensive identification of alternative back-splicing in human tissue transcriptomes, Nucleic Acids Research, Volume 48, Issue 4, 28 February 2020, Pages 1779–1789. [NAR]
Applications for circular oligonucleotides
Enzymatic synthesis of circular oligonucleotides
Extrachromosomal circular DNA is found in eukaryotic cells
Synthesis of circular oligonucleotides
Synthetic long single-stranded and circular DNA
What are circular oligonucleotides?
---...---
Bio-Synthesis provides circular oligonucletides of various lengths.
Bio-Synthesis provides a full spectrum of oligonucleotide and peptide synthesis including bio-conjugation services and high quality custom oligonucleotide modification services, back-bone modifications, conjugation to fatty acids and lipids, cholesterol, tocopherol, peptides as well as biotinylation by direct solid-phase chemical synthesis.
Enzyme-assisted approaches are also available to obtain artificially modified oligonucleotides, such as BNA antisense oligonucleotides, mRNAs or siRNAs.
Backbone modifications of base, sugar and internucleotide linkages are also possible.
---...---