The RAS gene family is one of the most studied and best characterized cancer-related genes (Arrington et al. 2012). KRAS is one of the three RAS isoforms that is the most frequently altered or mutated gene. KRAS mutations occur in 17 to 25% of all cancers. The Kristen Rat Sarcoma Virus and Murine Sarcoma Virus were co-discovered in 1982 by Chang and coworkers, and Der and Cooper (Jančík et al. 2010). The Harvey and Kirsten murine sarcoma viruses each encode a structurally and functionally related 21-kilodalton protein called p21. Ultimately the p21 protein was found to be the transforming protein of each virus. KRAS belongs to a group of small GTP-binding proteins also known as the RAS superfamily of RAS-like GTPases.
Figure 1: Molecular model of H-ras p21: The crystal structure for the H-ras oncogene protein 21 in complex with a slowly hydrolyzing GTP analog GppNp was determined at a resolution of 1.35 Å in 1990 (Pai et al. 1990). The binding site of the nucleotide and the magnesium could be revealed in high detail. This high resolution model allowed Pai et al. to propose a mechanism for GTP hydrolysis.
The KRAS gene, or Ki-ras2 Kirsten rat sarcoma viral oncogene homolog, is an oncogene encoding a small guanosine triphosphatase (GTPase) transducer protein called KRAS. The normal form of human c-Ras has also been called KRAS or KRAS2 or “Kristen Rat Sarcoma Viral oncogene homolog” or “Kristen Murine Sarcoma Virus 2 homolog.” The mammalian KRAS gene belongs to the ras gene family found on chromosomes 12p12.1 encoding a small GTPase. A single amino acid substitution or mutation results in a transforming protein.
In its natural, non-mutated or unchanged form the KRAS gene encodes for a protein called KRAS. KRAS is involved in cell signaling pathways that control cell growth, cell maturation, and cell death or apoptosis. The mutated or changed versions of the KRAS gene have been found in some types of cancer, including non-small cell lung cancer (NSCLC), colorectal cancer, and pancreatic cancer.
If a patient knows that his tumor has a wild-type or mutated KRAS gene, an optimal cancer treatment can be devised.
KRAS signaling
KRAS is one of the front-line sensors that initiate activation of an array of signaling molecules transmitting transducting signals from the cell surface to the nucleus. Transduction signaling pathways affect cell differentiation, growth, chemotaxis, and apoptosis. Signal transduction occurs in a cell when a molecule such as a hormone binds to a receptor on the cell membrane and initiates a set of chemical reactions inside the cell.
Many human tumors contain Ras mutations characteristically mutated at codons 12 or 61 but more rarely at 13. Most Gly12/Gln61 mutations are the chemical reason for the oncogenicity (the capability) of inducing tumor formation. The mutation prevents the RAS form of the protein from being switched off. The oncogenic Ras mutants remain constitutively activated and contribute to the abnormal growth of tissue of tumor cells. Missense mutations in RAS proteins alter the cells balance of GDP and GTP binding and thereby the active state of the protein. This can either occur by reducing GTP hydrolysis or by increasing the rate of GTP loading.
KRAS is frequently mutated in human cancers. The KRAS gene encodes for two distinct protein forms KRAS4A and KRAS4B through alternative splicing. The common mutations of KRAS produce mutant forms of KRAS4A and KRAS4B.
Mutations in the KRAS codon 12 are present in approximately 90% of ductal adenocarcinomas and in undifferentiated carcinomas of the pancreas. In colorectal cancer, KRAS gene mutations are correlated with increased proliferation and spontaneous apoptosis. Approximately 30 to 50 % of colorectal tumors have mutated KRAS genes.
Mutations in the KRAS oncogene are typically heterozygous. The ratio of the mutant allele to wild-type allele can be balanced or unbalanced. Sometimes, but relatively rare, the mutant allele can become dominant. This can occur either through deletion of the wild-type allele or copy number gain of the mutant allele. A mutation distribution observed in patients with KRAS-mutated pancreatic adenocarcinomas and undifferentiated pancreatic carcinomas is shown in table 1.
When the mutant becomes dominant, it is called “mutant allele-specific imbalance” or MASI. Cancers that have KRAS MASI appear to behave aggressively as is the case in lung and colon adenocarcinomas (cancers).
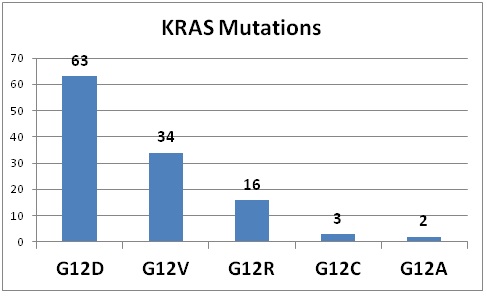
Table 1: Observed mutation distribution in patients with KRAS-mutated pancreatic adenocarcinomas and undifferentiated pancreatic carcinomas. (Source: Krasinskas et al. 2013.; Uniprot: http://www.uniprot.org/uniprot/P01116).
Glossary
Reference
Arrington AK, Heinrich EL, Lee W, et al. Prognostic and Predictive Roles ofKRAS Mutation in Colorectal Cancer. International Journal of Molecular Sciences. 2012;13(10):12153-12168. doi:10.3390/ijms131012153. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3497263/
Ahmadian MR, Zor T, Vogt D, et al. Guanosine triphosphatase stimulation of oncogenic Ras mutants. Proceedings of the National Academy of Sciences of the United States of America. 1999;96(12):7065-7070.
Chang EH, Gonda MA, Ellis RW, Scolnick EM, Lowy DR. Human genome contains four genes homologous to transforming genes of Harvey and Kirsten murine sarcoma viruses. Proceedings of the National Academy of Sciences of the United States of America. 1982;79(16):4848-4852. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC346782/.
Der, C. J., & Cooper, G. M. (1983). Altered gene products are associated with activation of cellular rasK genes in human lung and colon carcinomas. Cell, 32(1), 201-208. DOI: 10.1016/0092-8674(83)90510-X, http://www.sciencedirect.com/science/article/pii/009286748390510X
Hagis, K.M.; KRAS Alleles: The Devil Is in the Detail. Trends in Cancer, October 2017, 3, 10, 686-697. http://www.cell.com/trends/cancer/pdf/S2405-8033(17)30163-2.pdf
Krasinskas, Alyssa M, Moser, A James, Saka, Burcu, Adsay, N Volkan, Chiosea, Simion I; KRAS mutant allele-specific imbalance is associated with worse prognosis in pancreatic cancer and progression to undifferentiated carcinoma of the pancreas. Mod. Pathol. 2013/10/ 26 / 10 /1346 – 1354. http://dx.doi.org/10.1038/modpathol.2013.71
Kauke MJ, Traxlmayr MW, Parker JA, et al. An engineered protein antagonist of K-Ras/B-Raf interaction. Scientific Reports. 2017;7:5831. doi:10.1038/s41598-017-05889-7.
Liu X, Jakubowski M, Hunt JL.; KRAS gene mutation in colorectal cancer is correlated with increased proliferation and spontaneous apoptosis. Am J Clin Pathol. 2011 Feb;135(2):245-52. doi: 10.1309/AJCP7FO2VAXIVSTP.
Pai EF, Krengel U, Petsko GA, Goody RS, Kabsch W, Wittinghofer A. Refined crystal structure of the triphosphate conformation of H-ras p21 at 1.35 A resolution: implications for the mechanism of GTP hydrolysis. The EMBO Journal. 1990;9(8):2351-2359. https://www.ncbi.nlm.nih.gov/pubmed/2196171