The 5’-cap of eukaryotic mRNA is required for cellular function. The 5’-cap consist of an inverted 7-methylguanosine connected to the rest of the eukaryotic mRNA via a 5’-5’-triphosphate bridge (m7G cap or cap0). The cap, called cap0, acts as a quality control for correct mRNA processing. Cap0 contributes to the stability of eukaryotic mRNA, splicing, nuclear export, initiation of translation, and mRNA decay. Direct interaction molecules of the cap are the cap binding complex (CBC) in the nucleus needed for nuclear export and the eukaryotic translation initiation factor 4E (eIF4E) in the cytoplasm needed for cap-dependent translation.
Capped RNA is a marker for the innate immune system to distinguish triphosphorylated viral RNAs from cellular RNAs. The cytosolic receptor RIG-I mediates the antiviral response when activated by short single and double-stranded tri-phosphorylated RNAs and RNAs lacking the 2’-OH methylation at the first nucleotide (cap1). In addition to cap0 and cap1, other cap structures exist. Additional methyl groups can often be found at the second nucleotide (cap2).
Uncapped transcripts do not represent eukaryotic mRNAs therefore the preparation of correctly capped RNA is essential for synthetic mRNA used for the development of mRNA-based vaccines or RNA therapeutics. Altering the cap structure potentially increases mRNA stability and translational efficiency.
RNA polymerase II transcripts containing a protective 5',5'-triphosphate-linked 7-methylguanosine (m7G) cap can be decapped by enzymes like DCP2 initiating RNA decay. In the 5′–3′ mRNA decay pathway, poly(A) is shorted followed by cleavage of the mRNA cap. This reaction exposes the mRNA body to 5′–3′ exonucleases.
The capping of siRNAs increases their stability.
Wei et al., in 2013, used 1,2-bis(maleimido)ethane (BME) for the capping of hairpin RNAs to produce a capped siRNA called RhpRNA with a good serum and thermal stability. Teh protein Dicer was able to cleave these capped structures. RNA interference (RNAi) experiments showed that RhpRNA was highly efficient at RNAi with an IC50 value of 6 pM.
Adding a peptidyl cap to oligonucleotides stabilizes oligonucleotide base pairs.
Egetenmeyer & Richert, in 2011, identified a phosphodiester‐linked sequence of the residues of L‐prolinol, glycine, and oxolinic acid, dubbed ogOA, as a 5′‐cap that stabilizes any of the four canonical base pairs investigated, with ΔTm values of up to +13.1 °C for an oligonucleotide octamer.
NAD capping and Decapping
As reviewed by Megerditch Kiledjian, in 2018, eukaryotic cells contain nicotinamide adenine dinucleotide (NAD+)-capped RNAs. The NAD+ cap is added by transcriptional initiation with NAD+ in the place of ATP, and also by a novel NAD+ capping mechanism. The 5′-end NAD+ cap promotes the rapid decay of RNA. The decay appears to be mediated at least in part by the DXO family of proteins.
Recent reports showed that Saccharomyces cerevisiae and mammalian cells also contain mRNAs carrying a novel nicotinamide adenine dinucleotide (NAD+) cap at their 5′-end. The presence of an NAD+ cap on mRNA appears to be a new mechanism for controlling gene expression through nucleotide metabolite-directed mRNA turnover. The NAD+ cap targets RNA for rapid decay in mammalian cells through the DXO decapping enzyme by removing intact NAD+ from RNA in a process called ‘deNADding’. The DXO/Rai1 enzymes can eliminate most of the incomplete and non-canonical NAD caps through their decapping, deNADding and pyrophosphohydrolase activities as reviewed recently by Doamekpor et al. in 2020.
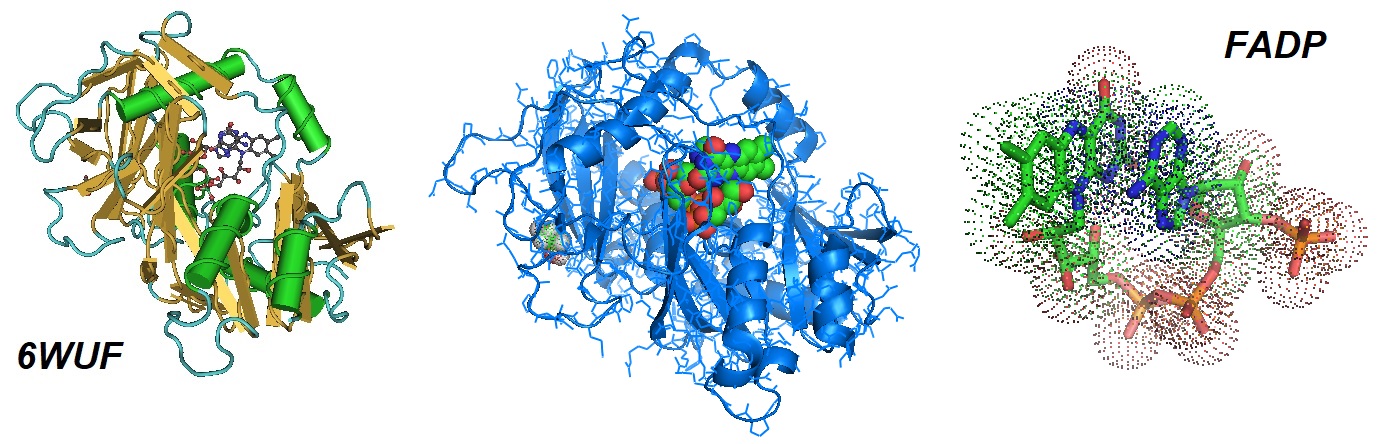
Figure 1: Structural model of the complex of mouse DXO wih 3’-FADP. In eukaryotes, the DXO/Rai1 enzymes can eliminate most of the incomplete and non-canonical NAD caps through their decapping, deNADding and pyrophosphohydrolase activities. According to Doamekpor et al., FAD caps are also present on short RNAs (with less than ∼200 nucleotides) in human cells. The absence of DXO stabilizes the RNA (Doamekpor et al. 2020).
Wu et al., in 2019, reported the structure of human NUDT12 and showed that it is a cytosolic NAD-RNA decapping enzyme. Each monomer of the active NUDT12 homodimers contributes to the two functional catalytic pockets. The ∼600-kDa dodecamer complex between bleomycin hydrolase (BLMH) and NUDT12 revealed that BLMH localizes NUDT12 to a few discrete cytoplasmic granules that are distinct from P-bodies. These two proteins, when artificially tethered to a reporter RNA, downregulate gene expression in vivo. The loss of Nudt12 resulted in a significant upregulation of circadian clock transcripts in the mouse liver. The study revealed a physiological role for NUDT12 in the cytosolic surveillance of NAD-RNAs.
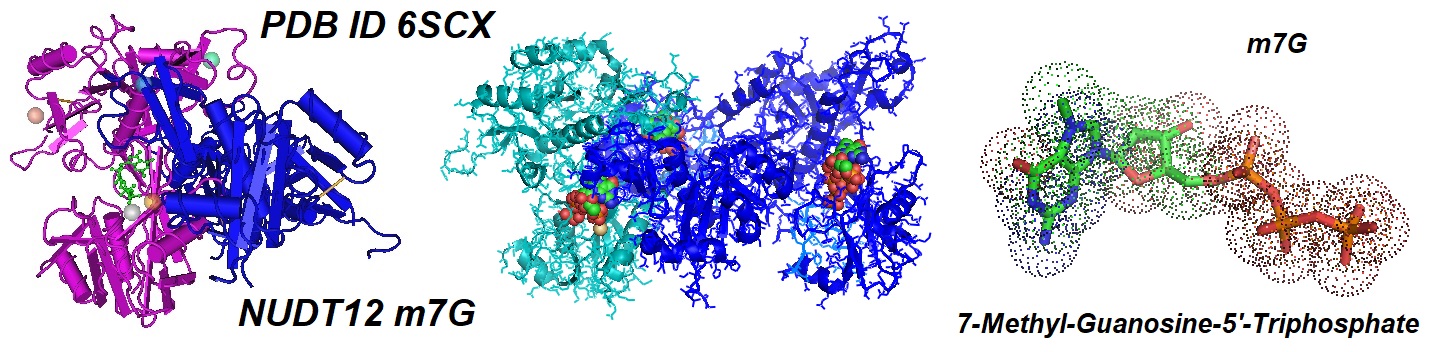
Figure 2: Crystal structure of the catalytic domain of human peroxisomal NADH pyrophosphatase NUDT12 in complex with 7-methyl-guanosine-5'-triphosphate.
Reference
Doamekpor, Selom, Grudzien-Nogalska, Ewa, Mlynarska-Cieslak, Agnieszka, Kowalska, Joanna, Kiledjian, Megerditch, Tong, Liang; 2020/05/06. DXO/Rai1 enzymes remove 5'-end FAD and dephospho-CoA caps on RNAs. Nucleic acids research 48, 10.1093/nar/gkaa297.. PDB ID 6WUF. [PMC]
Egetenmeyer, S.; Richert,; A 5′‐Cap for DNA Probes Binding RNA Target Strands. C. Chem. – Eur. J. 2011, 17, 11813–11827. [PubMed]
Megerditch Kiledjian; Eukaryotic RNA 5′-End NAD+ Capping and DeNADding. TICB; Volume 28, Issue 6, June 2018, Pages 454-464. [PMC]
Wei, L.; Cao, L.; Xi, Z. Highly Potent and Stable Capped siRNAs with Picomolar Activity for RNA Interference. Angew. Chem., Int. Ed. 2013, 52, 6501–6503.
Wu H, Li L, Chen KM, Homolka D, Gos P, Fleury-Olela F, McCarthy AA, Pillai RS. Decapping Enzyme NUDT12 Partners with BLMH for Cytoplasmic Surveillance of NAD-Capped RNAs. Cell Rep. 2019 Dec 24;29(13):4422-4434.e13. [PubMed]
---...---