Structured small RNAs natural, labeled or modified with nonstandard nucleotides can all be custom synthesized. These types of RNA are very useful and of high interest for structural and functional investigations.
Using modern chemical and biochemical methods, a diverse collection of RNA molecules, short, medium, and long can be prepared using solid-phase chemical synthesis or enzymatic synthesis or a combination of both synthesis methods.
Tracking RNA with fluorophores in vivo allows studying their complex cellular functions. The design of specific labels and labeling methods allows RNA painting as well as the study of various RNA species such as messenger RNA (mRNA), ribosomal RNA (rRNA), transfer RNA (tRNA), interference RNA (RNAi), small interfering RNA (siRNA), small nucleolar RNA (snoRNA), long-non-coding RNA (lncRNA), and others.
Splint ligation method
Longer RNA sequence fragments or blocks can be synthesized using ligation methods in combination with solid phase or enzymatic synthesis methods. The splint ligation approach allows assembly of smaller RNAs into larger RNA molecules. RNA fragments either synthesized via chemical solid phase synthesis or in-vitro transcription (IVT) are starting materials. This method allows precise labeling of long RNAs with modified groups or stable and radioactive isotopes. Ligation is possible using chemical synthesis methods or enzymatic methods using DNA and RNA ligases. A molecular “splint” is used to bring the two substrates together. The use of a splint enhances the ligation efficiency.
DNA ligase-based ligation
Ligation with T4 DNA ligase appears to be the most widely used method. T4 DNA ligase is largely sequence independent, tolerates modifications, and has minimal ligase activities with RNAs that are not hybridized to the DNA splint. In this approach, segments are joined using T4 DNA ligase-mediated splinted ligation, of two or more RNA fragments. RNA segments to be ligated are hybridized to a complementary DNA strand. This DNA strand is called the “splint.” Hybridization creates a nicked duplex in which the 3’-hydroxyl group of one RNA substrate is placed proximal to the 5’-monophosphate group of the other substrate.
Figure 1: T4 DNA or T4 RNA ligase ligation.
Sets of RNA fragments containing modified nucleotides can also be used. RNA fragments hybridize with the help of a complementary DNA splint forming a ternary ligation-competent-complex (LCC). This complex is turned over by the DNA ligase.
For highly structured RNAs, long splints are used to overcome the low propensity for hybridization and to significantly improve ligation efficiencies.
RNA ligase-based ligation
RNA ligases can also be used with similar strategies. However, T4 RNA ligase prefers ligation of single-stranded residues. In addition, the efficiency differs greatly between different RNA systems and efficiencies can vary between 40 to 80%.
Figure 2: Structural model of a nicked 5’-adenylated nucleic acid duplex containing a 2’-deoxyribonucleotide at the nick. The figure was created using the structural data from the T4 RNA ligase 2 nicked nucleic acid duplex, PDB 2HVS.
Chemical ligation
Chemical ligation is also possible. The phosphodiester bond linkage between two RNA substrates can be formed either by activating the phosphomonoester group using a reactive imidazolide or by using a condensing reagent such as cyanogen bromide. A disadvantage of chemical ligation is that it can also result in the creation of a 2’-5’ phosphodiester linkage, together with the desired 3’-5’ phosphodiester linkages. A purification step may be needed to produce a clean product.
Labeling RNA by hybridization
Labeled oligonucleotides are useful for the visualization of RNA using fluorescence resonance energy transfer (FRET). In this approach long target RNA recognize labeled oligonucleotides through Watson-Crick base-pairing. For FRET to work, two short labeled oligonucleotides are hybridized to two regions of a target RNA. The two modified oligonucleotides contain donor and acceptor fluorophores. The hybridization sites should be located such that the fluorophores are in the range of efficient energy transfer. To probe RNA tertiary structures the two hybridization sites can be located distantly in the primary sequence but should be located such that FRET can occur in the final tertiary structure. Also, oligonucleotides labeled with biotin are useful for single-molecule FRET. Synthetic labeled short DNA or RNA probes can be custom designed thereby allowing for a flexible and simple approach. It is important to design the probes such that they only anneal with non-functional regions of the target RNA to avoid steric hindrance during hybridization. To avoid false positive signals, correct experimental conditions need to be selected to avoid dissociation of non-covalent probes.
Figure 3: Hybridization based RNA labeling.
Aptamer based labeling method
Aptamers are oligonucleotides with sequences optimized for specific binding to a molecular target of interest. Aptamers have been made to specifically bind to fluorogenic dyes such as malachite green, Hoechst dye, fluorescein, and others. Also, RNA-binding proteins (RBPs) fused to probes, such as a green fluorescing protein, can be used to introduce probes into RNA molecules. For example, the phage MS2 coat protein that bind to hairpins in the 3’-untranslated region can be used to track the localization and dynamics of RNA. Unfortunately, GFP-RBP conjugates also emit fluorescence in the unbound form. This can interfere with the detection of target RNAs.
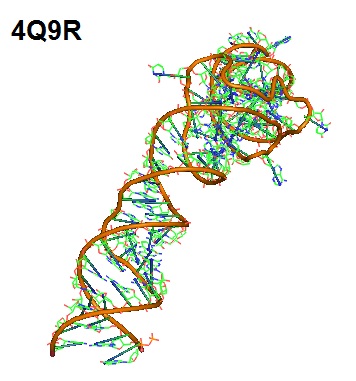
Figure 4: Aptamer Spinach. Spinach is an in vitro-selected RNA aptamer that binds a green fluorescein protein (GFP)-like ligand to activate its green fluorescence. Spinach is an RNA analog of GFP. Spinach binds the phenolate form of 3,5-difluoro-4-hydroxy-benzylidene imidazolinone (DFHBI) and selectively activates its fluorescence.
End-Labeling of Oligonucleotides
Several strategies for labeling the ends of oligonucleotides have been developed.
5’-end labeling using T4 PNK
This strategy makes use of the ability of bacteriophage T4 polynucleotide kinase (T4 PNK) to transfer a phosphate to the 5’-end of RNA or DNA oligonucleotides. If ATP is substituted with the ATP analog adenosine 5’-[γ-thio]triphosphate, the product of the transfer reaction is a phosphorylated oligonucleotide with a reactive sulfur at the 5’-end (1st reaction step). Incubation of the oligonucleotide with a haloacetamide derivative allows the addition of a chemical tag (2nd reaction step).
For example, 5-(iodoacetamido)fluorescein (5-IAF) can be used for conjugation of the fluorophore to the 5’-end of DNA and RNA oligonucleotides. Other compounds with different chemical properties can be used as well. However, for each reagent set used, optimization of the reaction will be necessary.
Figure 5: Attachment of a phosphorothioate to the 5’-terminal nucleotide.
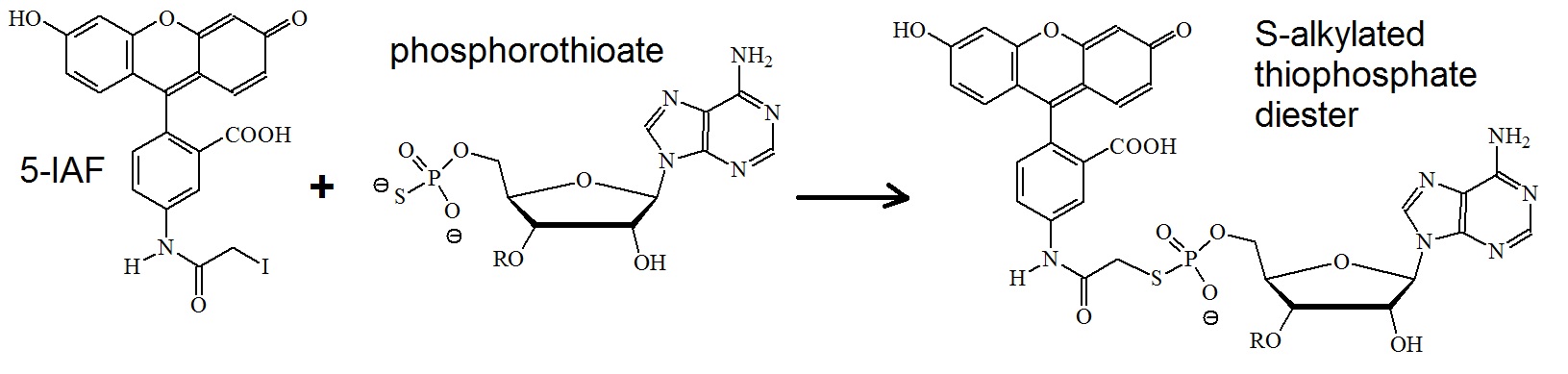
Figure 6: Reaction of 5-IAF with phosphorothioates. An S-alkylated thiophosphate diester is formed.
3’-end labeling using sodium periodate
This strategy makes also use of a two step reaction. First, sodium periodate oxidizes the 3’-terminal end of the ribose sugar. A reactive aldehyde is formed. Second, the oxidized sugar is conjugated to an aldehyde-reactive chemical tag. Often fluorescein 5-thio-semicarbazide (FITSC) is used. The periodate oxidation reaction requires the presence of vicinal hydroxyl groups. Therefore the reaction is specific for RNA and only modifies the 3’-terminal ribose.
Step 1: Formation of reactive aldehydes on 3’-end.
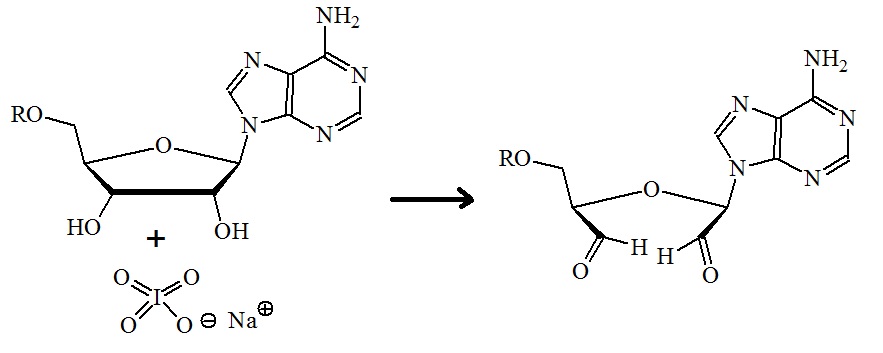
Figure 7: Oxidation of the diols on the 3’-terminal sugar. Sodium periodate is used for the oxidation to form reactive aldehydes. (For mechanism see Loudon M.; Organic chemistry. 5th edition. 2009, 506-507).
Step 2: Formation of a semicarbazone linkage between a fluorophore and an oligonucleotide.
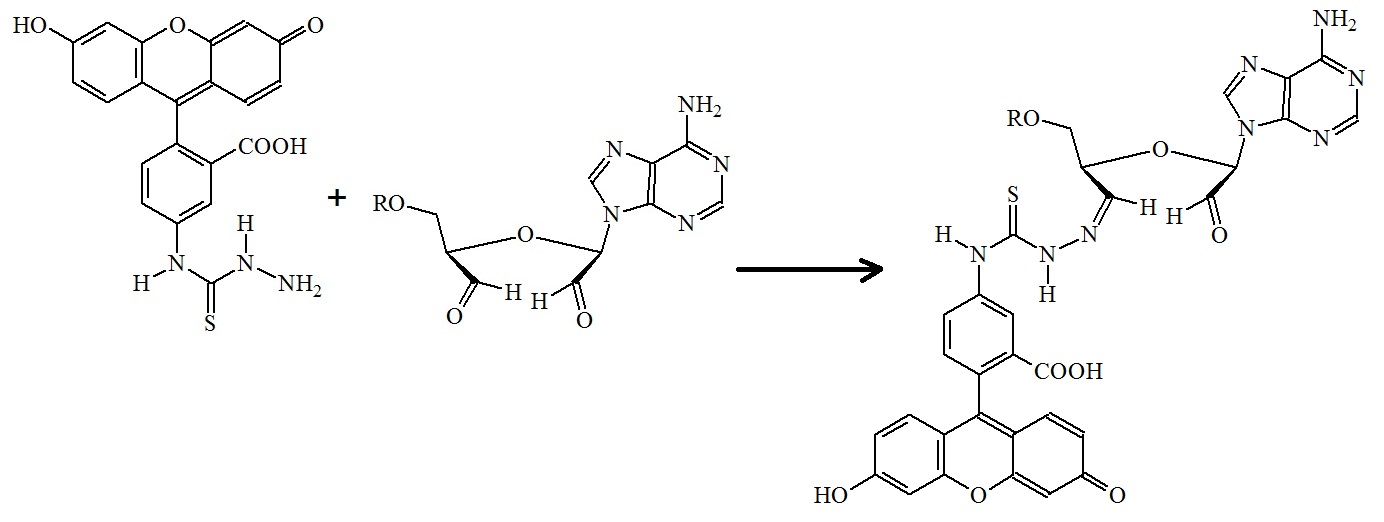
Figure 8: Formation of fluorescein 5-thiosemicarbazone. Fluorescein 5-thiosemicarbazide is incubated with the oxidized 3’-terminal sugar.
A variety of aldehyde-reactive fluorescent compounds are now commercially available for selective labeling of 3’-end RNA. Also, oligonucleotides can be conjugated to biotin using (+)-biotinamidohexanoic acid hydrazide (BACH), or to a solid support using adipic-acid dihydrazide-agarose.
Reference
Eastberg JH, Pelletier J, Stoddard BL. Recognition of DNA substrates by T4 bacteriophage polynucleotide kinase. Nucleic Acids Research. 2004;32(2):653-660. doi:10.1093/nar/gkh212. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC373337/
Huang H, Suslov NB, Li N-S, et al. A G-Quadruplex-Containing RNA Activates Fluorescence in a GFP-Like Fluorophore. Nature chemical biology. 2014;10(8):686-691. doi:10.1038/nchembio.1561.
KURSCHAT WC, MÜLLER J, WOMBACHER R, HELM M. Optimizing splinted ligation of highly structured small RNAs. RNA. 2005;11(12):1909-1914. doi:10.1261/rna.2170705.
Liu, Y., Sousa, R., and Wang, Y.-X.; Specific labeling: An effective tool to explore the RNA world. Bioessays 38: 192-200.
Maroney, Patricia A; Chamnongpol, Sangpen; Souret, Frédéric; Nilsen, Timothy W.; Direct detection of small RNAs using splinted ligation. Nature Protocols 3, 279 – 287 (2008). http://dx.doi.org/10.1038/nprot.2007.530
Nandakumar, Jayakrishnan et al.; RNA Ligase Structures Reveal the Basis for RNA Specificity and Conformational Changes that Drive Ligation Forward. Cell , Volume 127 , Issue 1 , 71 – 84.
---...---