Cellular Genomes
Cellular genomes are very vulnerable to damage during chromosomal DNA replication can cause mutations as a result of the incorporation of incorrect bases during DNA replication. For the maintenance of genomic integrity, cells have several mechanisms to repair these damages. For example, the reversal of chemical reactions that caused DNA breakage and the removal of damaged bases followed by their replacement with newly synthesized DNA. Cells also have other means to cope with damaged DNA.
The protein proliferating cell nuclear antigen (PCNA) initially identified as an antigen in the nuclei of DNA synthesizing cells, is a DNA clamp essential for DNA repair. PCNA is a homotrimer protein complex that encircles DNA. Bound to DNA, it functions as a scaffold for the recruitment of proteins needed for the replication of DNA, remodeling of chromatin as well as for epigenetic modifications. Many of proteins required for DNA repair interact with PCNA via the PCNA-interacting motif called PCNA-interacting peptide box (PIP) or the AlkB homolog 2 PCNA interacting motif (APIM). PCNA can function as a sliding clamp during DNA synthesis, a polymerase switch factor, or as a recruitment factor. Interactions of PCNA with various proteins together with post-translational modifications of PCNA control the regulation of the cell cycle and chromatid cohesion.
Wu et al. recently identify the E3 ubiquitin ligase TRAIP (TRAF-interacting protein) as a factor located at active and stressed replication forks. TRAIP directly interacts with PCNA via a conserved PCNA-interacting peptide (PIP) box motif. TRAIP promotes ATR-dependent checkpoint signaling in human cells via the generation of replication protein A (RPA)-bound single-stranded DNA regions. Replication stress requires its E3 ligase activity, which is potentiated by the PIP box. Wu et al. report that the loss of TRAIP leads to chromosomal instability and decreased cell survival after replication stress. TRAIP appears to be a PCNA-binding ubiquitin ligase protecting genome integrity after difficult DNA replication.
RPA is the major eukaryotic ssDNA-binding protein with essential roles in genome maintenance. RPA binds to ssDNA through multiple dynamic modes. RPA can alternate between different binding ways modifying ssDNA structures. However, many of these interactions remain unknown.
TRAIP and DNA repair
Cellular genomes are very vulnerable to damage during chromosomal DNA replication. Alterations of genomic DNA can cause mutations as a result of the incorporation of incorrect bases during DNA replication. For the maintenance of genomic integrity, cells have several mechanisms to repair these damages. For example, the reversal of chemical reactions that caused DNA breakage and the removal of damaged bases followed by their replacement with newly synthesized DNA. Cells also have other means to cope with damaged DNA.
The protein proliferating cell nuclear antigen (PCNA), initially identified as an antigen in the nuclei of DNA synthesizing cells, is a DNA clamp essential for DNA repair. PCNA is a homotrimer protein complex that encircles DNA. Bound to DNA, it functions as a scaffold for the recruitment of proteins needed for the replication of DNA, remodeling of chromatin as well as for epigenetic modifications. Many of proteins required for DNA repair interact with PCNA via the PCNA-interacting motif called PCNA-interacting peptide box (PIP) or the AlkB homolog 2 PCNA interacting motif (APIM). PCNA can function as a sliding clamp during DNA synthesis, a polymerase switch factor, or as a recruitment factor. Interactions of PCNA with various proteins together with post-translational modifications of PCNA control the regulation of the cell cycle and chromatid cohesion.
Wu et al. recently identify the E3 ubiquitin ligase TRAIP (TRAF-interacting protein) as a factor located at active and stressed replication forks. TRAIP directly interacts with PCNA via a conserved PCNA-interacting peptide (PIP) box motif. TRAIP promotes ATR-dependent checkpoint signaling in human cells via the generation of replication protein A (RPA)-bound single-stranded DNA regions. Replication stress requires its E3 ligase activity, which is potentiated by the PIP box. Wu et al. report that the loss of TRAIP leads to chromosomal instability and decreased cell survival after replication stress. TRAIP appears to be a PCNA-binding ubiquitin ligase protecting genome integrity after difficult DNA replication.
RPA is the major eukaryotic ssDNA-binding protein with essential roles in genome maintenance. RPA binds to ssDNA through multiple dynamic modes. RPA can alternate between different binding ways modifying ssDNA structures. However, many of these interactions remain unknown.
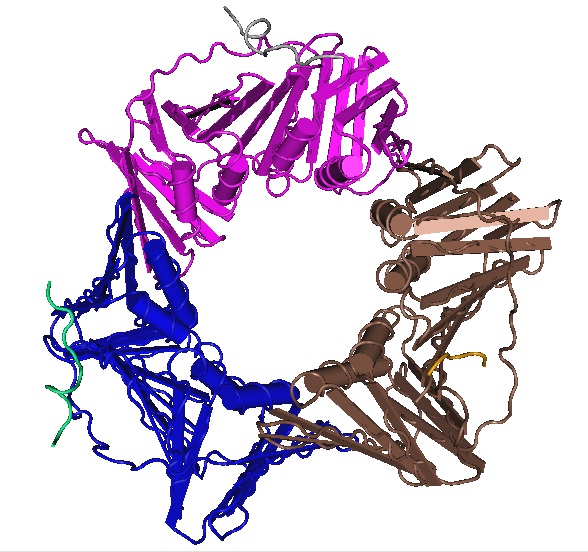
Crystal structure of human PCNA in complex with a TRAIP peptide.
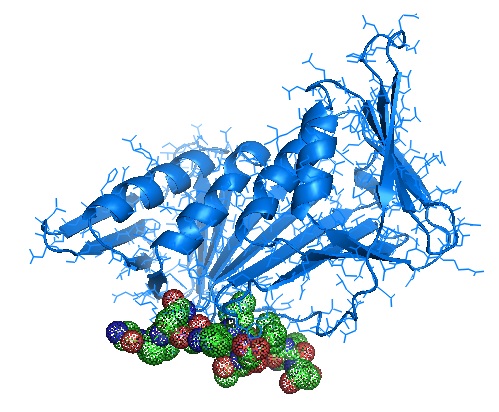
Monomer structure of PCNA in complex with Ala-phe-gln-ala-lys-leu-asp-thr-phe-leu-trp-ser and Ala-gly-ala-gly-ala [4ZTD].
Sequence alignment of the peptide sequence with the protein sequence shows that the interacting peptide is located at the C-terminal end of E3 ubiquitin-protein ligase TRAIP.
E3 ubiquitin-protein ligase TRAIP [Homo sapiens]
>NP_005870.2 E3 ubiquitin-protein ligase TRAIP [Homo sapiens]
MPIRALCTICSDFFDHSRDVAAIHCGHTFHLQCLIQWFETAPSRTCPQCRIQVGKRTIINKLFFDLAQEE
ENVLDAEFLKNELDNVRAQLSQKDKEKRDSQVIIDTLRDTLEERNATVVSLQQALGKAEMLCSTLKKQMK
YLEQQQDETKQAQEEARRLRSKMKTMEQIELLLQSQRPEVEEMIRDMGVGQSAVEQLAVYCVSLKKEYEN
LKEARKASGEVADKLRKDLFSSRSKLQTVYSELDQAKLELKSAQKDLQSADKEIMSLKKKLTMLQETLNL
PPVASETVDRLVLESPAPVEVNLKLRRPSFRDDIDLNATFDVDTPPARPSSSQHGYYEKLCLEKSHSPIQ
DVPKKICKGPRKESQLSLGGQSCAGEPDEELVGAFPIFVRNAILGQKQPKRPRSESSCSKDVVRTGFDGL
GGRTKFIQPTDTVMIRPLPVKPKTKVKQRVRVKTVPSLFQAKLDTFLWS
Earlier in 2017, Klattenhoff et al. showed that the protein Nei endonuclease VIII-like 3 (NEIL3) is also localized at DNA double-strand break (DSB) sites during oxidative DNA damage and replication stress.
Loss of NEIL3 significantly increased spontaneous replication-associated DSBs and recruitment of replication protein A (RPA). In contrast, we observed a marked decrease in Rad51 on nascent DNA strands at the replication fork, suggesting that HR-dependent repair is compromised in NEIL3-deficient cells. Interestingly, NEIL3-deficient cells were sensitive to ataxia–telangiectasia and Rad3 related protein (ATR) inhibitor alone or in combination with PARP1 inhibitor. This study elucidates the mechanism by which NEIL3 is critical to overcome oxidative and replication-associated genotoxic stress.
Reference
DNA repair: https://www.ncbi.nlm.nih.gov/books/NBK9900/
https://en.wikipedia.org/wiki/Proliferating_cell_nuclear_antigen
https://www.ncbi.nlm.nih.gov/books/NBK9900/
Deeba N. Syed, Mohammad Imran Khan, Maria Shabbir, Hasan Mukhtar; MicroRNAs in Skin Response to UV Radiation. Current Drug Targets Volume 14 , Issue 10 , 2013. DOI : 10.2174/13894501113149990184. [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3985496/]
Klattenhoff AW, Thakur M, Chu CS, Ray D, Habib SL, Kidane D. Loss of NEIL3 DNA glycosylase markedly increases replication associated double strand breaks and enhances sensitivity to ATR inhibitor in glioblastoma cells. Oncotarget. 2017 Dec 4;8(68):112942-112958. doi: 10.18632/oncotarget.22896. PMID: 29348879; PMCID: PMC5762564. [https://www.ncbi.nlm.nih.gov/pubmed/29348879]
Strzalka W, Ziemienowicz A. Proliferating cell nuclear antigen (PCNA): a key factor in DNA replication and cell cycle regulation. Ann Bot. 2011 May;107(7):1127-40. doi: 10.1093/aob/mcq243. Epub 2010 Dec 17. PMID: 21169293; PMCID: PMC3091797. [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3091797/]
Qing-Man Wang, Yan-Tao Yang, Yi-Ran Wang, Bo Gao, Xuguang Xi and Xi-Miao Hou; Human Replication protein A induces dynamic changes in single-stranded DNA and RNA structures. The Journal of Biological Chemistry294, 13915-13927. [http://www.jbc.org/content/early/2019/07/26/jbc.RA119.009737.abstract]
Wu RA, Semlow DR, Kamimae-Lanning AN, Kochenova OV, Chistol G, Hodskinson MR, Amunugama R, Sparks JL, Wang M, Deng L, Mimoso CA, Low E, Patel KJ, Walter JC. TRAIP is a master regulator of DNA interstrand crosslink repair. Nature. 2019 Mar;567(7747):267-272. doi: 10.1038/s41586-019-1002-0. Epub 2019 Mar 6. PMID: 30842657; PMCID: PMC6417926. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6417926.