Coronaviruses are known to cross the species barrier and infect humans. The recent outbreak of SARS-CoV-2 (COVID-19) is a vivid example. The outbreaks of SARS in 2003 and, more recently, Middle-East respiratory syndrome (MERS) already demonstrated the lethality of coronaviruses (CoVs).
Since 2003, much progress has been made in understanding aspects of the virus’ life cycle. The CoV envelope (E) protein is a small, integral membrane protein taking part in virus assembly, budding envelope formation, and pathogenesis. Recent biochemical studies increased our knowledge of E protein’s structure, structural motifs, its function as an ion-channeling viroprotein, as well as its interaction with other coronavirus proteins and host cell proteins.
Coronaviral genomes encode four major structural proteins required to produce a structural complete viral particle. The four major structural proteins are the spike or surface protein (S), nucleocapsid (N), membrane (M) protein, and the envelope (E) protein. Some C0Vs do not need the E protein to form a complete, infectious virion.
The E protein is the smallest of the major structural proteins. During the replication cycle, the virus expresses the E protein in high abundance inside the cell. However, only a small portion is incorporated into the virion envelope. Most of the protein is localized at the site of intercellular trafficking, at the endoplasmic reticulum (ER), the Golgi, and the Endoplasmic reticulum Golgi intermediate compartment (ERGIC). The E protein participates in viral assembly and budding. Coronaviruses without the E protein appear to mature much slower than CoVs in which the E protein is present.
The E protein of coronaviruses is a conserved, short, integral membrane protein of 76 to 109 amino acids, ranging in size from 8.4 to 12 kDa. Figure 1 shows the alignment of E protein sequences for MERS, SARS1, SARS-CoV-2, sequences from solved protein structures 5X29, 2MM4, 5XES, and the self-assembly peptide TVYVYSRVK.

Figure 1: Alignment of CoV E proteins for MERS, SARS and SARS CoV-2, and the C-terminal peptide TVYVYSRVK known to self-assemble.
The E protein has a short, hydrophilic amino terminus of 7 to 12 amino acids, followed by a large hydrophobic transmembrane domain (TMD) of 25 amino acids, and a hydrophilic carboxyl terminus (Figure 2). The hydrophobic TMD region has at least one amphipatic α-helix that can oligomerize to form an ion-conductive pore of membranes. Sequence analysis of both, E and S protein for SARS and SARS CoV-2, revealed a triple cysteine motif located after the transmembrane region of the E protein (LCAYCN) and in the C-terminal end of the S protein (SCGSCCK). Wu et al., in 2003, suggested that these two motifs could serve as a structural basis for the association between the E and S protein. However, this may need to be experimentally verified. Synthetic peptides spanning the motif could serve as tools for the study of these motifs.

Figure 2: Amino acid sequences and domains of the E protein from MERS, SARS and SARS-CoV-2 virus. The E protein contains three domains, the amino (N)-terminal domain, the transmembrane domain (TMD), and the carboxy (C)-terminal domain. The putative interaction peptide and the C-terminal peptide known to self-assemble are also indicated within the sequence.
Li et al., in 2014, reported that the E protein in the severe acute respiratory syndrome virus has only one transmembrane (TM) domain in micelles and that the predicted β-coil-β motif forms a short membrane-bound α-helix connected by a disordered loop to the TM domain. More recently, Surya et al., in 2018, reported the structural model of the SARS coronavirus E channel in lyso-myristoyl phosphatidylglycerol (LMPG) micelles (Figure 3).
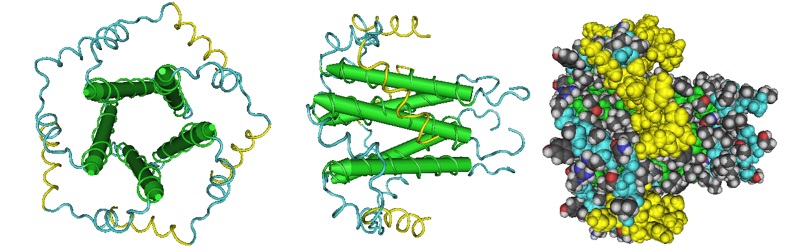
Figure 3: NMR model of the SARS coronavirus E channel in LMPG micelles (Surya et al.; PCB ID 5X29). The model of the pentamer was optained by docking the monomeric form of the protein using the software HADDOCK 2.2. In LMPG micelles, the C-terminal tail of SARS-Cov E protein is α-helical.
Reference
Li Y, Surya W, Claudine S, Torres J. Structure of a conserved Golgi complex-targeting signal in coronavirus envelope proteins. J Biol Chem. 2014 May 2;289(18):12535-49. [PMC]
Schoeman, D., Fielding, B.C. Coronavirus envelope protein: current knowledge. Virol J 16, 69 (2019).
Surya W, Li Y, Torres J. Structural model of the SARS coronavirus E channel in LMPG micelles. Biochim Biophys Acta Biomembr. 2018 Jun;1860(6):1309-1317. [PMC]
Wu Q, Zhang Y, Lü H, Wang J, He X, Liu Y, et al. The E protein is a multifunctional membrane protein of SARS-CoV. Genomics, Proteomics & Bioinformatics. 2003;1(2):131–44. [Article]
Bio-Synthesis Inc. is pleased to offer a large variety of DNA/RNA oligonucleotides and peptides, modified or unmodified, conjugates, as well as stapled peptides, for a number of research applications, including COVID 19 testing, analysis and vaccine development!
---...---