What are lncRNAs and lincRNAs?
Long noncoding RNAs, or lncRNAs, are recently discovered RNA molecules. lncRNAs represent a class of long transcribed RNA molecules that are longer than 200 nucleotides. lncRNA transcripts do not code for proteins.
Long intergenic noncoding RNAs, or lincRNAs, are long RNA transcripts that have been identified in mammalian genomes by bioinformatics analysis of transcriptomic data. Thousands of lincRNAs are now known, however, many of their functions are still unknown. lincRNAs appear to contribute to the control of cell differentiation and maintenance of cell identity.
Recent discoveries increasingly indicate that only a small percentage, approximately 7%, of disease-associated single nucleotide polymorphisms (SNPs) are located in protein-coding regions, whereas the remaining 93% are located in gene regulatory regions or in intergenic regions. Therefore, our understanding of how genetic variations control the expression of non-coding RNAs tissue-specific will have far reaching implications for how diseases will need to be treated in the future.
Advances made in high-throughput RNA sequencing technologies have provided scientists an expanded view of the complexity of a genome, and it has now become clear that most of the human genome is transcribed to produce not only protein-coding transcripts, but also large numbers of non-coding RNAs (ncRNAs) of different size. By now many types of RNA have been well characterized, such as short ncRNAs, microRNAs, small interfering RNAs, and piwi-interacting RNAs.
However, it was discovered that the large intergenic non-coding RNAs (lincRNAs) make up most of the long ncRNAs. LincRNAs are non-coding transcripts of more than 200 nucleotides long; they have an exon-intron-exon structure, similar to protein-coding genes, but do not encompass open-reading frames and do not code for proteins. More than 8,000 lincRNAs have been descript recently and it is thought that lincRNAs are the largest subclass of RNAs, originating from the non-coding transcriptome in humans.
According to Cabili et al. (2011), "large intergenic non-coding RNAs (lincRNAs) are the largest class of non-coding RNA molecules in the human genome. Many genome-wide association studies (GWAS) have mapped disease-associated genetic variants (SNPs) to, or in, the vicinity of such lincRNA regions. At this point in time it is not clear how these SNPs can affect the disease.” Cabili et al. tested whether SNPs were also associated with the lincRNA expression levels in five different human primary tissues. They observed that there is a strong genotype-lincRNA expression correlation that is tissue-dependent. Many of the observed lincRNA cis-eQTLs are disease- or trait-associated SNPs. Their results suggested that lincRNA-eQTLs represent a novel link between non-coding SNPs and the expression of protein-coding genes, which can be exploited to understand the process of gene-regulation through lincRNAs in more detail. Note that expression quantitative trait loci (eQTLs) are genomic loci that regulate expression levels of mRNAs or protein.
Long non-coding RNAs are non-protein coding sequence transcripts that contain more than 200 nucleotides. Their size distinguishes lncRNAs from small regulatory RNAs such as microRNAs (miRNAs), short interfering RNAs (siRNAs), Piwi-interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs), short hairpin RNA (shRNA), and other short RNAs. Since the central dogma of gene expression is that DNA is transcribed into messenger RNAs that serve as the templates for protein synthesis, lncRNAs where considered in the past as “junk” DNA or the “dark matter” of DNA. The discovery of extensive transcription of large RNA molecules that do not code for proteins provided a new perspective on the roles of RNA in gene regulation.
New findings from the study of multiple model systems suggest that lncRNAs form extensive networks of ribonucleoprotein (RNP) complexes with numerous chromatin regulators. These chromatin regulators target these enzymatic activities to the appropriate locations in the genome. Apparently lncRNAs can function as modular scaffolds to specify higher-order organization in RNP complexes and in chromatin states. Recent research in human transcriptome analysis shows that protein-coding sequences only account for a small portion of the genome transcripts. The majority of the human genome transcripts are non-coding RNAs now called long-non-coding RNAs (lncRNAs). Chen et al. in 2013 describe a long-non-coding RNA (lncRNA) and disease association database (LncRNADisease).
The database is publicly accessible at http://cmbi.bjmu.edu.cn/lncrnadisease.
Suggested Functions of lncRNAs
In recent years, a variety of regulatory paradigms for how long ncRNAs may function have been identified. LncRNAs may function by binding to DNA or RNA in a sequence specific manner or by binding to proteins. In contrast to miRNAs, lncRNAs appear not to operate by a common mode of action but can apparently regulate gene expression and protein synthesis in a number of ways.
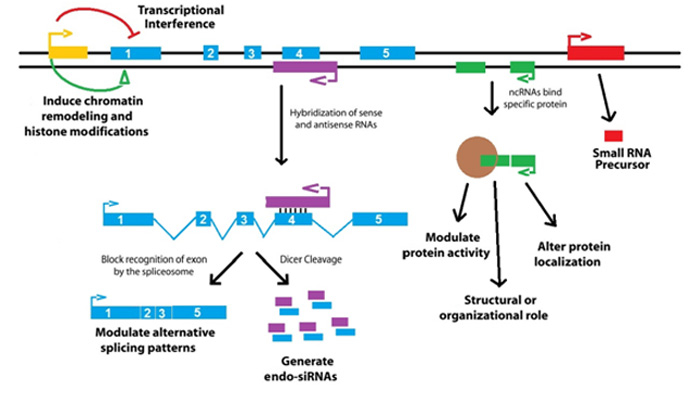
1. Transcription
Transcription from an upstream noncoding promoter (yellow) can negatively or positively affect expression of the downstream gene (blue) by inhibiting RNA polymerase II recruitment or inducing chromatin remodeling, respectively.
2. Antisense Transcripts
An antisense transcript (purple) is able to hybridize to the overlapping sense transcript (blue) and block recognition of the splice sites by the spliceosome. This results in an alternatively spliced transcript.
3. Hybridzation of sense to antisense transcripts
Hybridization of the sense and antisense transcripts can allow Dicer to generate endogenous siRNAs.
4. Binding to noncoding transcripts
Binding of the noncoding transcript (green) to specific protein partners can modulate the activity of the protein or serve as a structural component that allows a larger RNA–protein complex to form, or alter where the protein localizes in the cell.
5. LncRNA processing
Long ncRNAs (red) can be processed to yield small RNAs, such as miRNAs, piRNAs, and other less well-characterized classes of small transcripts.
Classification of lncRNA
lncRNAs can be classified into the following locus biotypes based on their location with respect to protein-coding genes:
|
1
|
Intergenic lncRNA
|
Intergenic lncRNAs are transcribed inter genetically from both strands.
|
|
2
|
Intronic lncRNA
|
Intronic lncRNAs are entirely transcribed from introns of protein-coding genes.
|
|
3
|
Sense lncRNA
|
Sense lncRNAs are transcribed from the sense strand of protein-coding genes and contain exons from protein-coding genes that overlap with part of protein-coding genes or cover the entire sequence of a protein-coding gene through an intron.
|
|
4
|
Antisense lncRNA
|
Antisense lncRNAs are transcribed from the antisense strand of the protein-coding genes that overlap with exonic or intronic regions, or cover the entire protein-coding sequence through an intron.
|
Furthermore, it needs to be pointed out that presently there is still an uncertainty between comparative results. However, it has become clear that lncRNAs are emerging as regulatory elements of embryonic pluripotency, differentiation, and patterning of the body axis as well as promoting developmental transitions.
Potential Functions and Molecular Mechanisms of lncRNAs
Wang and Chang in 2011 illustrated the construction of complex functions by using combinations of archetypical molecular mechanisms of lncRNAs.

Schematic diagram of the four archetypes of lncRNA mechanisms.
(Adapted from Wang, and Chang, Mol Cell. 2011).
|
Archetype I
|
As Signals: LncRNA expression can faithfully reflect the combinatorial actions of transcription factors (colored ovals) or signaling pathways to indicate gene regulation in space and time.
|
|
Archetype II
|
As Decoys: LncRNAs can titrate away transcription factors and other proteins away from chromatin, or titrate the protein factors into nuclear subdomains. A further example of decoys is a lncRNA decoy for miRNA target sites (not shown on schematic).
|
|
Archetype III
|
As Guides: LncRNAs can recruit chromatin modifying enzymes to target genes, either in cis (near the site of lncRNA production) or in trans to distant target genes.
|
|
Archetype IV
|
As scaffolds: LncRNAs can bring together multiple proteins to form ribonucleoprotein complexes. The lncRNA-RNP may act on chromatin as illustrated to affect histone modifications. In other instances, the lncRNA scaffold is structural and stabilizes nuclear structures or signaling complexes.
|
lncRNAs and disease and research
lncRNAs are now implicated in a variety of diseases. Recent studies have shown that lncRNAs are expressed differently in various types of cancer including leukemia, breast cancer, hepatocellular carcinoma, colon cancer, and prostate cancer. In other diseases, such as cardiovascular diseases, neurologicaldisorders and immune-mediated diseases, lncRNA appear to be deregulated.
Since lncRNA can be present in very low amounts, can overlap with coding transcripts on both strands, and are often only found in the nucleus, working with this type of RNA can be very challenging. The table below list differences and similarities of mRNAs, lncRNAs and lincRNAs.
A comparison of mRNAs, lncRNAs and lincRNAs
Advantages of BNA-enhanced research for ncRNA
Bridged nucleic acids (BNA) are artificial bicyclic oligonucleotides that contain a six-membered bridged structure with a “fixed” C3’-endo sugar puckering. The bridge is synthetically incorporated at the 2’, 4’-position of the ribose to afford a 2’, 4’-BNA monomer. BNAs are structurally rigid oligo-nucleotides with increased binding affinities and stability.
BNA monomers can be used for both primers and probes in real time quantitative polymerase chain reaction (RT-Q-PCR) assays. Compared to locked nucleic acids (LNAs), the substitution of DNA monomers with BNA monomers in oligonucleotides adds exceptional biological stability, resistance to nucleases and a significantly increased affinity to their complementary DNA targets.
In addition, short, high affinity, BNA-enhanced qPCR primers can enhance the detection of low abundant targets. Furthermore, the specific placement of BNA monomer within the oligonucleotide probe allows to adjust the melting temperature (Tm) of the probe which may be important for qPCR analysis of overlapping transcripts.
Applications of BNA Oligonucleotides for lncRNA and lincRNA Research
-
RNA isolation
-
Custom BNAs for qPCR assay development for mRNA and ncRNA
-
Custom BNA detection probes for ncRNAs
-
BNA long RNA Gapmer Antisense Oligonucleotides
-
Custom BNA Oligonucleotides
Ordering & Contact Information
- Please contact us for additional information or send an email to info@biosyn.com.
- You may also request an online quote.
- To contact us by phone, please call 1-972-420-8505 or Fax at 1-972-420-0442
- Orders may be placed using a purchase order (PO) or by credit card through our secure online ordering system.
- When using a credit card (
 ) it will be billed under "Bio-Synthesis, Inc."
) it will be billed under "Bio-Synthesis, Inc."
References
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, et al. (2011); Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25: 1915–1927. Access the most recent version at doi:10.1101/gad.17446611.
HUGO Gene Nomenclature Committee - HGNC: http://www.genenames.org/
Wang KC, Chang HY.; Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011 Sep 16;43(6):904-14. doi: 10.1016/j.molcel.2011.08.018.