Exogenous mRNA allows the development of mRNA-based therapeutics to treat a variety of diseases such as genetic disorders, cancer, infectious diseases, cardiovascular diseases, and vaccine development. Recent advances in chemical modifications of mRNA structures to synthesize modified mRNAs improved the stability and translational efficiency within cells when delivering exogenous mRNAs.
Recent examples for the use of exogenous mRNAs are the new vaccines against COVID-19 from Moderna, the COVID-19 vaccine (mRNA-1273), which is an mRNA vaccine against COVID-19 encoding for a prefusion stabilized form of the Spike (S) protein, and the Pfizer-BioNTech COVID-19 vaccine, also encoding the S protein of the virus.
Regulation of cellular gene expression occurs at the post-transcriptional level. Messenger RNA carries genetic information from the nucleus to the cytoplasm needed for the production of functional proteins. For a cell to function well, normal levels of mRNAs need to be maintained. mRNA is a single-stranded RNA transcribed from a DNA template sequence. mRNAs contain untranslated regions that control translation, degradation, and localization of the mRNA’s coding sequence.
RNA sequence regions of mRNAs include stem-loop structures, upstream initiation codons and open reading frames, internal ribosome entry sites, and various cis-acting elements that interact with RNA-binding proteins.
Typically, a mature eukaryotic mRNA consists of a 5’-methylguanosine (m7G) cap, a 5’-untranslated region (UTR), a coding region starting with the AUG codon, a 3’-UTR, and a polyadenylated A (poly(A)) tail. The Kozak sequence, GCCACCAUGG, in the 5’-UTR around the initiating methionine allows for sufficient ribosomal binding to initiate translation.
The 3’-UTR is also essential in regulating mRNAs' translation and stability and may contain miRNA binding sites. The poly(A) tail is a critical component for mRNA translation and degradation. Degradation of mRNAs starts from the 5'-cap structure if the poly(A) tail is shorter than 12 adenosine residues.
The stability and translation efficiency of exogenous mRNAs can be enhanced by many methods, including manipulations of UTRs, codon optimization, chemical modification, and the elongation of the poly(A) tail.
Transcription Control.
A variety of factors control or mediate transcription. Factors mediating transcription are transcription factors, RNA polymerase, a variety of cis-acting elements located in the DNA. Cis-acting elements include promotors, enhancers, silencers, and locus-control elements organized in a modular structure regulating pre-mRNA production.
Pre-mRNAs undergo several processing steps to become functional mRNAs. These are:
(i) Removal of introns.
(ii) Addition of a 7-methyl-guanylate (m7G) cap structure at the 5’-end of the first exon.
(ii) Addition of 100 to 250 adenine residues (the poly(A) tail) at the 3’-end of the last exon.
(iV) Cleavage of the primary transcript by endonucleolytic processing of the primary transcript.
9v) mRNA editing.
In eukaryotes, the result is a mature mRNA with a modular structure consisting of a 5'-untranslated region (5'-UTR), a coding region made up of triplet codons that each encode an amino acid, and a 3'-untranslated region (3'-UTR). Figure 1 shows these and other features of mRNAs.
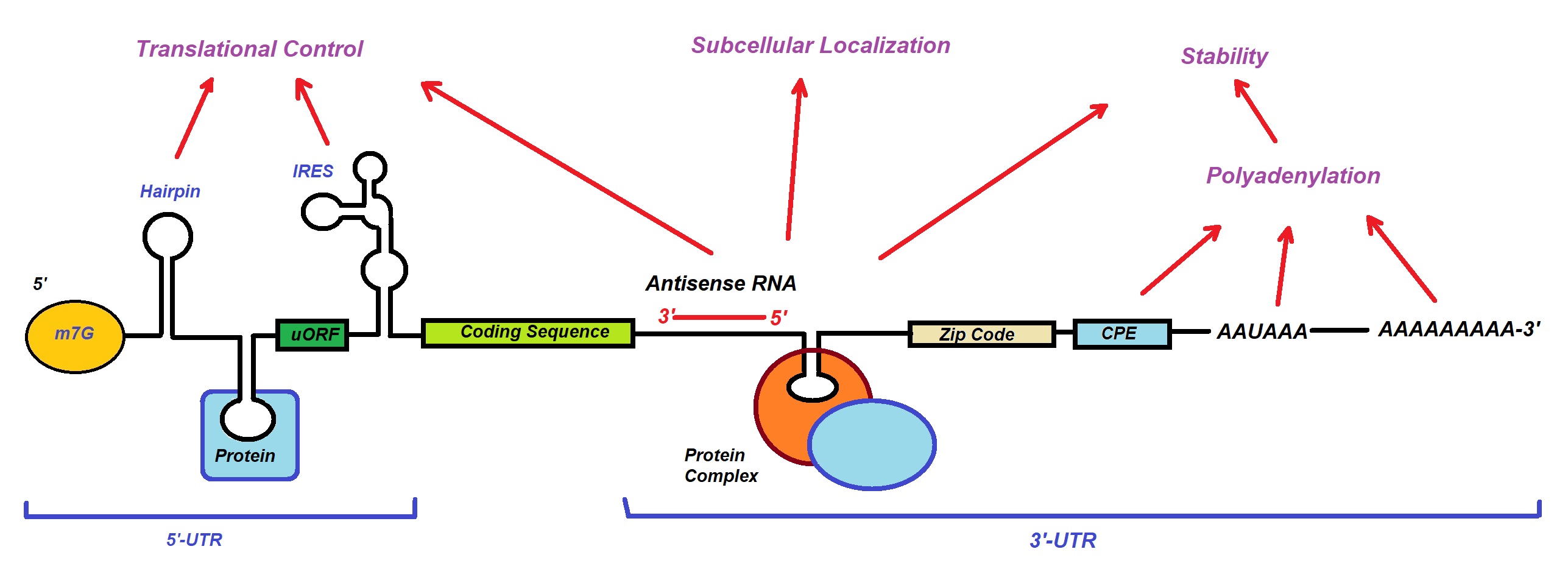
Figure 1: Model of a generic structure of a eukaryotic mRNA. Some post-transcriptional regulatory elements that affect gene expression are illustrated (Adapted from Mignone et al., 2020).
Special UTRs
Modified amino acid selenocysteine at UGA codons mediated by a conserved stem-loop structure in the 3’-UTR.
Nucleotide patterns or motifs located in the 5’-UTRs and 3’-UTRs can interact with specific RNA-binding proteins.
Interactions of sequence elements located in the UTRs and specific complementary non-coding RNAs play vital roles in regulation.
Repetitive elements such as CUG repeats in the 5’-UTR can bind to CUG-binding proteins. The transcription factor C/EBPβ is an example.
Also, the nuclear history of an mRNA can affect its fate in the cytoplasm.
Genomic sequence comparison revealed that the average length of 5’-UTRs ranges in size between 100 to 200 nucleotides and that it is roughly constant over diverse taxonomic classes.
The average length of 3’-UTRs can vary between 200 to 800 nucleotides between species.
Sometimes a single nucleotide in the 5’-UTR can initiate translation.
The G+C content in 5’-UTR sequences is greater than that of 3’-UTR sequences.
Genes localized in large GC-rich regions of a chromosome have shorter 5’-UTRs and 3’-UTRs than genes located in the GC-poor areas or isochores. Isochores are large DNA regions containing a high degree of guanine and cytosine (G-C and C-G).
Eukaryotic mRNAs contain several types of repeat sequences in the untranlated region such as short interspersed elements (SINEs; Alu elements), long interspersed elements (LINEs), minisatellites and microsatellites.
In humans, repeats are found in approximately 12% of 5’-UTRs and 36% of 3’-UTRs.
Reference
Chemical and enzymatic synthesis of mRNA and modified mRNA
COVID-19 Vaccines: Moderna, COVID-19 vaccine (mRNA-1273), and Pfizer-BioNTech COVID-19 vaccine
Li B, Zhang X, Dong Y. Nanoscale platforms for messenger RNA delivery. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2019 Mar;11(2):e1530. [PMC]
Mignone F, Gissi C, Liuni S, Pesole G. Untranslated regions of mRNAs. Genome Biol. 2002;3(3):REVIEWS0004. doi: 10.1186/gb-2002-3-3-reviews0004. Epub 2002 Feb 28. [PMC]
mRNA modifications
Pichon X, Wilson LA, Stoneley M, Bastide A, King HA, Somers J, Willis AE. RNA binding protein/RNA element interactions and the control of translation. Curr Protein Pept Sci. 2012 Jun;13(4):294-304. [PMC]
RNA capping: 5’-methylguanosine (m7G) cap
Nishat Sultana, Yoav Hadas, Mohammad Tofael Kabir Sharkar, Keerat Kaur, Ajit Magadum, Ann Anu Kurian, Nadia Hossain, Bremy Alburquerque, Sakib Ahmed, Elena Chepurko, and Lior Zangi; Optimization of 5' Untranslated Region of Modified mRNA for Use in Cardiac or Hepatic Ischemic Injury. Molecular Therapy: Methods & Clinical Development Vol. 17 June 2020. [link]
---...---
Bio-Synthesis provides a full spectrum of high quality custom oligonucleotide modification services by direct solid-phase chemical synthesis or enzyme-assisted approaches to obtain artificially modified oligonucleotides containing backbone, base, sugar and internucleotide linkages including messenger RNA.
Bio-Synthesis specialize in complex oligonucleotide modifications using phosphodiester backbone, purine and pyrimidine heterocyclic bases, and sugar modified nucleotides such as our patented 3rd generation Bridged Nucleic Acids.
Bio-Synthesis also provides synthetic capped RNA oligonucleotides.