MicroRNAs (miRNAs) are short RNA oligonucleotides with an average length of 22 nucleotides. miRNAs play essential roles in many biological processes, including cell development, proliferation, differentiation, and apoptosis. Thousands of miRNAs are known as regulatory molecules modulating gene expression at post-transcriptional levels. Correlating differently expressed miRNAs in tissues with cellular processes may allow diagnostics of various diseases. Hence, the detection of miRNA can provide valuable diagnostic and prognostic data. However, because of their size, measurements of miRNAs are challenging.
In 2016, Aviñó et al. designed and synthesized modified parallel tail-clamps to detect miRNA sequences responsible for tumor suppression via triplex formation. The researchers used parallel clamps targeting a seven homo-pyrimidine sequence in miRNA-145 to test if they form triple helices (Triplex; See Figure 1). The research group selected 8-aminoG instead of 8-aminoA for improved triplex stability because of the small polypyrimidine tracks present in miRNAs.
A pH 5 was selected to test the complex's thermal stability between parallel clamps and miRNA-145. This pH range is known to be optimal for parallel-triplex formation. A surface plasmon resonance (SPR) biosensor allowed the detection of miRNA-145, forming a triplex with parallel clamps.
Triplex froming oligonucleotides (TFOs) are a class of DNA oligonucleotides capable of binding to the major groove of duplex DNA froming a triplex helix. Triplex forming oligonucleotides, for example, containing bridged nucleic acids (BNAs) can also be used to inhibit expression of specific genes.
Triplex formed
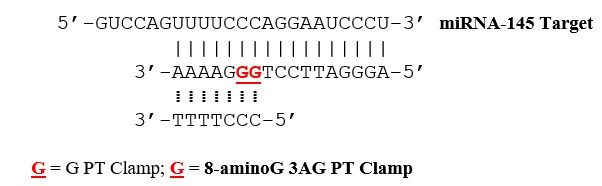
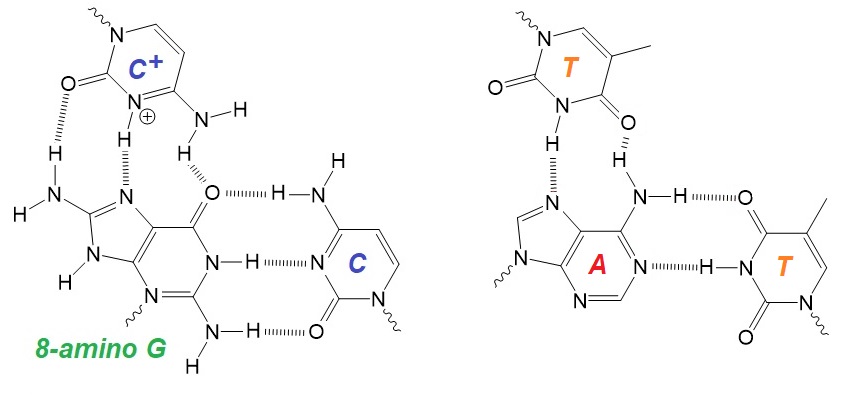
Figure 1: Tail-clamps and complementary sequences forming a miRNA-145 triplex structure. The triads C+-8-aminoG-C and T-A-T are shown as well.
Avino et al. found that parallel clamps with 8-aminoguanine form the most stable triplex structures with targeted miRNA. SPR biosensor analysis demonstrated that the target miRNA and the 8-aminoguanine tail-clamp formed a stable triplex structure. The researchers proposed to use this approach for label-free and reliable detection of miRNA signatures needed for diagnostic purposes.
Reference
Aviñó, A., Huertas, C. S., Lechuga, L. M., Eritja, R.; Sensitive and label-free detection of miRNA-145 by triplex formation. Anal. Bioanal. Chem., 408(3), 885-893 (2016). [PubMed]
---...---