In Drosophila, A’-form RNA helices drive microtubule-based mRNA transport
A cell uses microtubule-based mRNA transport to restrict protein expression to a specific location within the cell. The exact location of proteins in cells defines cell polarity, axis determination, and neuronal function. RNA localization coupled to translational repression creates structural and functional asymmetry within cells. Cytoskeletal elements such as microtubules and actin filaments associated with mechanoenzymes mediate the transport of a significant fraction of transcripts. The best-studied model systems of RNA localization are the oocyte and the early embryo in Drosophila melanogaster.
In eukaryotes, asymmetric localization of mRNAs plays widespread roles in protein targeting important for cellular processes such as patterning of embryonic axes, polarized cell functions, and synaptic plasticity. To understand the structural basis of recognition of cis-acting mRNA location signals by motor complexes Bullock et al. in 2010 used NMR spectroscopy for the determination of the tertiary structure of an RNA element responsible for mRNA transport.
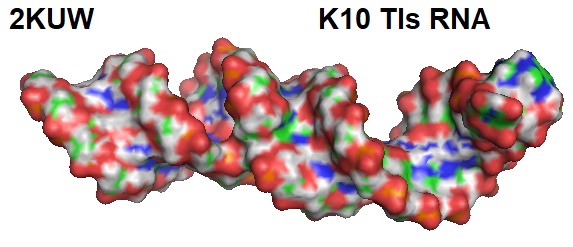
Figure 1: Surface model of the solution structure of K10 TLS RNA.
According to Bullock et al. the K10 TLS RNA adopts a stem-loop structure capped by an octanucleotide loop (5′-A(18)UUAAUUC(25)-3′) displaying a compact fold. The helical part of the TLS can consist of three regions: an upper helix composed of seven Watson-Crick A-U or U-A base pairs, a middle helix of three Watson-Crick base pairs, flanked at each end by single nucleotide bulges on the 3′ side, and a lower helix consisting of a G-U and six Watson-Crick base pairs. The two unpaired bases adopt different orientations relative to the helices, and the base moiety of C33 resides in the major groove maintains the helical twist between the adjacent base pairs. The base of A37 is stacked in between the middle and the lower helix and increases the helical twist between the adjacent base pairs.
Knowing the structure of this RNA molecule adds to our understanding of cis-acting mRNA localization signals by molecular motor complexes. The K10 transport localization element is a regulatory element considing of 44 nucleotides. This RNA structure is responsible for the transport and anterior localization of k10 mRNA in the oocyte in which it establishes dorsoventral polarity.
For the study of molecular principles of mRNA localization, important developmental transcripts are delivered to the minus-ends of microtubules during early Drosophila development. The localization process is dynein-dependent and microinjection of in vitro synthesized fluorescent transcripts allow their study. Already several minus-end-directed transport signals have been mapped in Drosophila mRNAs. These RNA signals are all predicted to adopt stem-loop structures containing ~ 40-65 nucleotides (nt). However, they do not necessarily share primary sequences or any obvious RNA motifs. However, at present, it is not clear which features within mRNAs are recognized by the transport machinery. More studies will be needed to elucidate the molecular mechanisms involved in mRNA localization.
Reference
Bullock SL, Ringel I, Ish-Horowicz D, Lukavsky PJ. A'-form RNA helices are required for cytoplasmic mRNA transport in Drosophila. Nat Struct Mol Biol. 2010;17(6):703-9.
---...---