The quality of the synthesis method and instruments used for the synthesis influences the final yield!
The theoretical maximal amount of product possibly produced from any particular synthesis is determined by the quality of the synthesis itself. Most syntheses of oligonucleotides are now done using automated solid phase oligonucleotide synthesizers. Here, the coupling efficiency of the synthesis method is very important. The coupling effciency determines how long an oligonucleotide can be syntheisized as well as the quality of the final product.
At Bio-Synthesis we do our best to optimize this efficiency using modified programs as well as paying special attention to the preparation of all reagents employed. However, a drop in coupling efficiency of just a few percentage points can lower the actual yield dramatically.
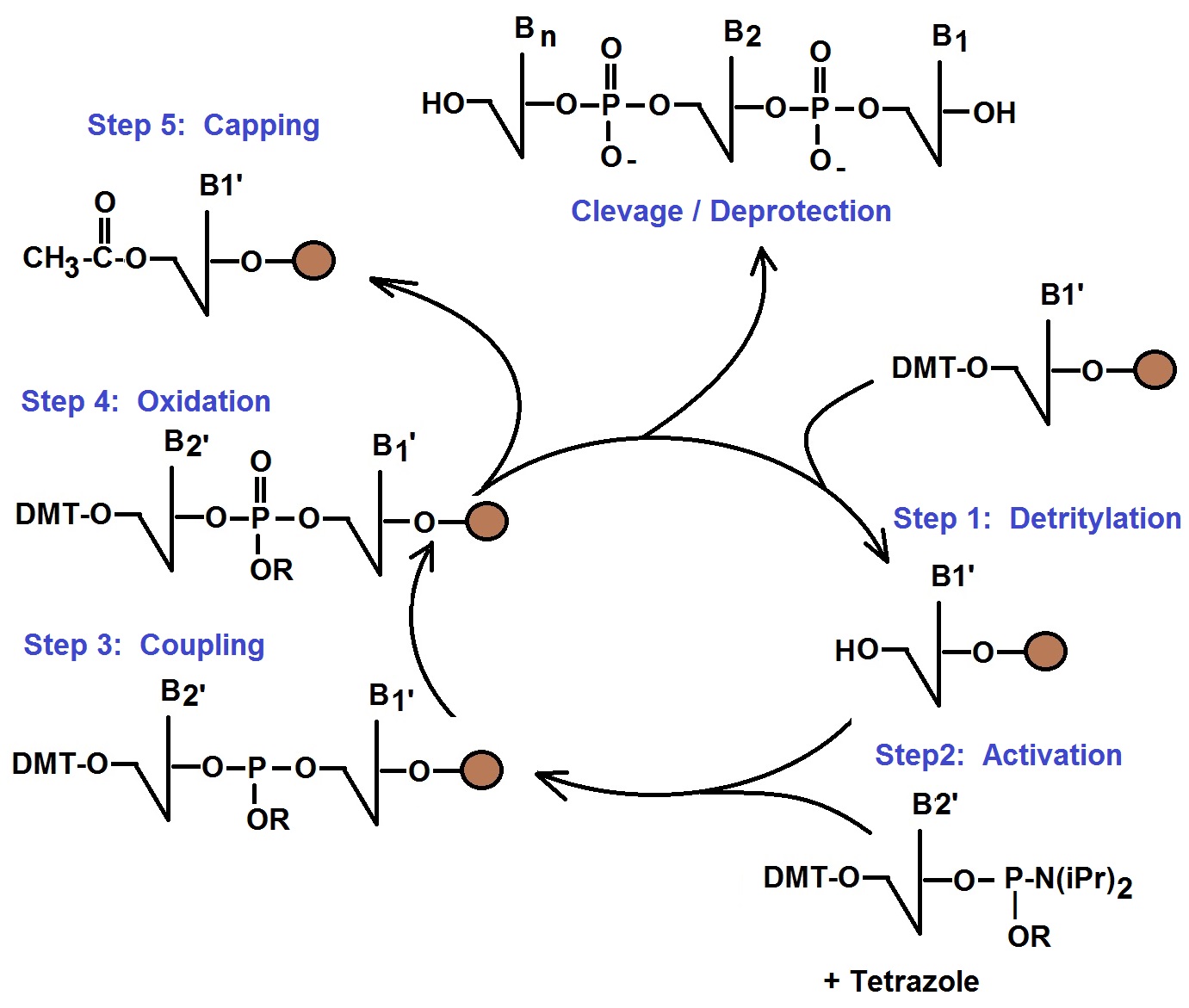
Phosphoramidite Method of Oligonucleotide Synthesis
Calculating Theoretical Yields
The quality of a polymer synthesis as well as the performance of a oligonucleotide synthsizer can be evaluated using the average coupling effciency data. As can bee seen from plotting the oligomer lenght versus theoretical overall yield with respect to the coupling effciency, the higher the coupling effcieny of a synthesis is, the longer oliomer can be synthesized.
Plot of Oligomer Length versus Theoretical Overall Yield
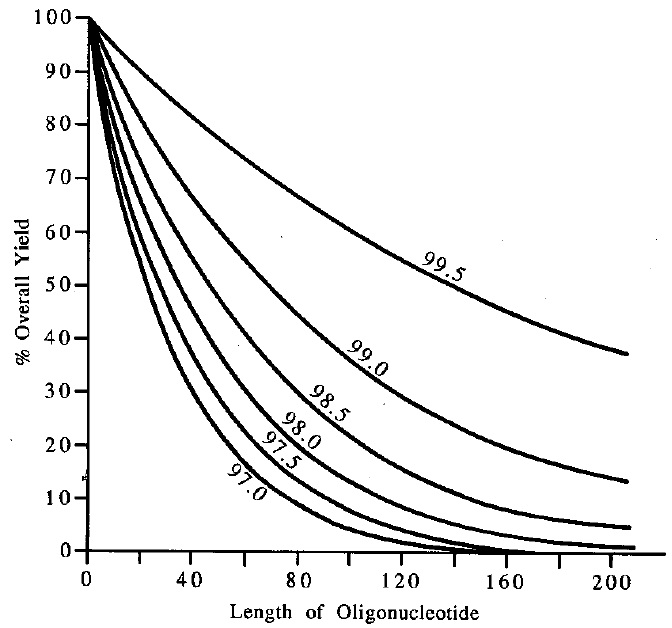
Basically two methods for the determination of the average coupling effciency for a synthesis are commonly used. A high coupling efficiency is very important for the synthesis of long oligonucleotides.
Trityl Method
One, is still used today in a Expedite synthesizer, is based on monitoring the orange trityl fractions. These are collected from each coupling cycle and are plotted for each cycle number.
HPLC Method
The second method is based on the analysis of chromatographic data generated using HPLC.
However, ultimately the analysis of the final product will varify the quality of the product.
Theoretical Yield
The theoretical yield can be calculate using the formula: Yieldth = XY
where "x" is the average coupling efficiency and "y" is the number of coupling.
For example, a synthesis of a 30mer (which requires 29 couplings) with an average coupling efficiency of 99%, theoretically yields 75% of product (0.9929). However, the same synthesis at a 98% efficiency will have a maximum yield of only 55%. That one percent difference in effciency costs nearly half of the material. Consider the difficulty of making a 70 mer. Even at 99%, the best one could hope for is 50% yield. At 98% it becomes an much lower yield of 25%.
It is very difficult to maintain every synthesizer every day at an operational efficiency of 99% or greater, despite claims to the contrary. Not only does the instrument have to be finely tuned to operate at 99%+, but other factors have to be perfect, such as the moisture content in the acetonitrile and phosphoramidite quality. The weather plays a role, as well. Extremely humid days will adversely affect the quality of a synthesis by making near complete water removal almost impossible, despite using rigorous anhydrous chemistry techniques.
Modified reagents often have poor coupling efficiencies for a variety of reasons. It is not unusual for a reagent to have a coupling efficiency as low as 90%.
Some modified oligonucleotides are prepared by conjugation to an amine, thiol or carboxyl functionality. This reaction is affected by many factors as well, including the quality and age of reagent, the sequence of oligonucleotide and the quality of the buffers. Reactions can range from 10% to nearly quantitative. A mediocre conjugation can have a considerable effect on the final product yield.
At BSI we do our best to control the quality of all reagents through our ISO9001:2015 program, as well as using well-established SOPs and trained personnel. Despite our best efforts, from time to time some reactions will still progress poorly due to sequence effects.
Deprotection
After synthesis, the oligonucleotide is deprotected with the treatment of a base. For simple oligonucleotides this presents little difficulty, as long as the deprotecting reagents are fresh. However, particular modifications require deprotecting conditions so mild that the bases are not fully deprotected. In most cases, some damage occurs to the bases themselves, but particularly to dye labeled compounds. These side reactions will all result in a lower yield.
Purification
Depending on the quality of the synthesis, purification can be the step where the most of the yield is lost. A high quality synthesis will have only a moderate amount of impurities to remove, allowing a larger cut of the product peak. Moderate and poor syntheses will have more contaminating fragments that will crowd into the product peak, requiring a tighter cut to obtain an acceptable purity. Regardless of the quality of the synthesis, the overall process of purification will result in a lower overall yield. Upwards of 50% of the theoretical yield will be consumed in many preparations for a wide variety of reasons. Oligonucleotides that are partially deprotected and those containing degraded or damaged dyes often share enough properties with the product to make purification difficult. In some cases, the modifications on an oligonucleotide will cause the product to co-elute with shorter fragments, which is the case with unprotected primary amine modified oligonucleotides. The requested purity makes a considerable impact on the yield. The difference in delivered product between a final purity of 90% and 95% can be several fold.
What should I expect from my 1 µmole scale modified oligonucleotide order?
Here is a hypothetical synthesis to illustrate how yield is affected. We will return to the example of the 30mer described above, but add the following: it has a 5'-dye and a 3'-quencher attached. This hypothetical dye is only available as a succinimidyl ester, therefore requiring post-synthesis conjugation to a 5'-amino labeled oligonucleotide. The 3'-quencher in this case is a support bound reagent from which the synthesis begins, requiring no further chemistry. The oligonucleotide is further modified with three modified bases, each of which in this example couple at 93%. The amino linker couples with an efficiency of 95%.
Therefore (0.9926)(0.933)(0.95) = 0.59
The deprotection of the oligonucleotide always results in a small amount of undeprotected bases, undesired modifications, and in this case, some degradation of the quencher. It also involves transfers and filtrations that invariably result in most of the loss. Overall, a loss of another 25% of potential product due to chemical modifications, incomplete deprotections and manual manipulation is not unusual.
Our yield is now at 44%, or 0.44 µmole.
The protecting group on the amino group is left on to improve the purification of the amino labeled intermediate prior to conjugation. Without it, the amino labeled oligonucleotide will elute with the failure sequences on HPLC. It is common to lose 10% of this group prior to purification during deprotection and work-up, lowering our yield to 0.40 μmole.
The purification of this product will result in another loss of at least 25% of the product due to contamination with the before mentioned, undesired modified materials.
We are now at 0.30 μmole of intermediate.
The conjugation of a dye generally reacts at 70-90% unless there is a sequence or reagent issue. Let us assume an efficiency of 80%.
The yield is now at 0.24 μmole.
This material is now re-purified to remove unconjugated material, free dye, and any side products due to damaged dyes, resulting in a further loss of 20% of the product, reducing the product to 0.19 μmole.
This is followed by a series of manipulations to desalt the sample, convert it to the proper salt and precipitate it, and then remove aliquots for final analysis prior to deliver. Unfortunately, this series of steps is more costly than appreciated, resulting in an overall loss of another 20% of the product, reducing the final yield to 0.15 μmoles.
Given a 30mer with an ε of 345 (OD260 units)(mL) (μmole)-1 and a molecular weight of 10,500 g/m, or 10.5 mg/μmole, this final yield would correspond to a delivery of 52 OD260 units, or 1.6 mg, of product.
This yield is considered as an excellent yield for a highly modified compound like this. Yields of 20-40 OD260 units for compounds of this sort are much more common. The losses described can easily be compounded by many different factors, some completely out of our control (mostly construct/sequence issues) and some within our control (level of training, routine maintenance). A compound with high Guanosine content can cause the loss of at least half of the material if it has a secondary structure that makes purification difficult. Also, recall that a day with higher than usual humidity can result in the loss of a third of the product right from the onset of the synthesis by dropping coupling efficiency just one percentage point.
When should I request set quantity instead of synthesis scale?
Almost all large-scale synthesis (50mg and larger) are ordered by set quantity, usually in units of mass. Some customers order 10 mg samples, depending on us to decide on the proper starting scale. There are cases when ordering a set quantity for smaller amounts may make sense. Many diagnostic companies require guaranteed amounts of material delivered so that they can produce the allotted number of kits, even at small quantities. This will increase your cost considerably in some cases since we may have to prepare a larger scale to ensure the yield you requested.
However, it is best to indicate your desired final quantity needed. Bio-Synthesis can decide a scale of synthesis for you.
---...---