Horseradish Peroxidase, or HRP, allows the design and synthesis of molecular probes such as in situ hybridization probes to increase signal amplification. HRP, when conjugated to hybridization probes, increases signal amplification, for example, in Catalyzed Reporter Deposition-Fluorescent In Situ Hybridization (CARD-FISH) applications.
Historically, horseradish peroxidase is used as a reporter enzyme in diagnostics and histochemistry applications. Horseradish Peroxidase can be conjugated to oligonucleotides, peptides, proteins, or other molecules, for example, via a maleimide attachment to form a thioester bond.
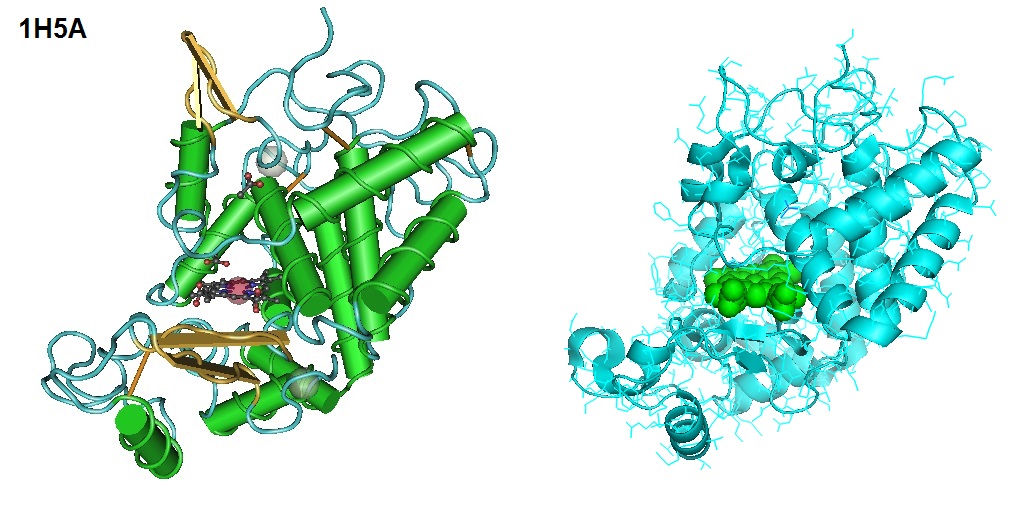
Horseradish Peroxidase is an enzymatically active glycoprotein. HRP is a globular molecule with an α-helical secondary structure and one short β-sheet region (Gajhede et al. 1997).
The HRP molecule is separated into a distal and a proximal region by an iron protoporphyrin IX cofactor. The iron protoporphyrin IX cofactor is known as the heme group. Heme is typically linked to HRP by a coordinate bond of the heme iron with a conserved His170 residue. In the presence of H2O2, HRP catalyzes an oxidation reaction that converts reduced, non-fluorescent or non-colorimetric substrates into oxidized, fluorescent or colorimetric molecules. Furthermore, HRP C1A contains nine Asn-X-Ser/Thr-X motives (X being any amino acid but Pro) which are potential N-glycosylation sites.
For conjugation reactions, surface lysine residues allow directed enzyme immobilization or conjugation to target compounds via covalent linkages. On the surface of HRP C1A, three out of six lysine residues were found accessible to chemical modifications. These are Lys174, Lys232, and Lys241 (O’Brien et al. 2001).
Reference
Ferrari, B. C., Tujula, N., Stoner, K., & Kjelleberg, S. (2006). Catalyzed Reporter Deposition-Fluorescence In Situ Hybridization Allows for Enrichment-Independent Detection of Microcolony-Forming Soil Bacteria. Applied and Environmental Microbiology, 72(1), 918–922. [PubMed]
Gajhede M, Schuller DJ, Henriksen A, Smith AT, Poulos TL. Crystal structure of horseradish peroxidase C at 2.15 A resolution. Nat Struct Biol. 1997;4:1032–8. [PubMed]
Krainer, F. W., & Glieder, A. (2015). An updated view on horseradish peroxidases: recombinant production and biotechnological applications. Applied Microbiology and Biotechnology, 99, 1611–1625. [PubMed]
O’Brien AM, O’Fágáin C, Nielsen PF, Welinder KG. Location of crosslinks in chemically stabilized horseradish peroxidase: implications for the design of crosslinks. Biotechnol Bioeng. 2001;76:277–84. [PubMed]