Yes, BSI provides custom designed site-specifically labeled oligonucleotides?
Site-specific labeled oligonucleotides are essential tools used as DNA probes or substrates for the characterization of DNA or RNA binding to proteins. For example, site-specific labeled oligonucleotides allow studying translocation and unwinding events of helicases. Assays are available or can be designed to determine equilibrium binding affinities for single-stranded DNA (ssDNA), ssRNA, as well as duplex and branched DNA and RNA substrates. Fluorophores used for the design of a variety of oligonucleotide substrates are usually cyanine 3 and 5 (Cy3, Cy5) dyes.
Furthermore, the intrinsic fluorescence quenching of the inherent fluorescence of native tryptophan (Trp) residues, when present in the protein, allows studying the orientation and position of a helicases when bound to DNA or RNA.
In addition, the iron-sulfur (FeS) clusters found in many helicases can serve as an endogenous quencher of Cy3 and Cy5 fluorescence. This phenomenon allows characterization of helicases containing the FeS cluster.
Well designed fluorescence probes and substrates enable the determination of the translocation rate by the helicase at the single-molecule level.
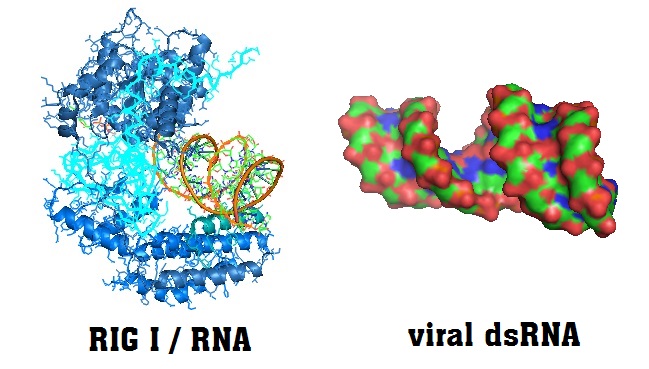
Structure of the RNA binding protein duck retinoic acid-inducible gene I (RIG I) RNA helicase.
RIG I triggers the cellular innate interferon (IFN) immune response upon detection of viral double-stranded RNA. As the structural analysis of the protein reveales, the helicase needs the cofactor adenosin-5’-diphosphate (ADP) and magnesium ions for the recognition of intracellular 5'-triphosphate double-stranded RNA (5'-ppp-dsRNA).
Reference
Kowalinski, Eva et al.; Structural Basis for the Activation of Innate Immune Pattern-Recognition Receptor RIG-I by Viral RNA. Cell , Volume 147 , Issue 2 , 423 – 435.
Robert A.Pugh, Masayoshi Honda, MariaSpies; Ensemble and single-molecule fluorescence-based assays to monitor DNA binding, translocation, and unwinding by iron–sulfur cluster containing helicases. Methods Volume 51, Issue 3, July 2010, Pages 313-321.
Flowers S1, Biswas EE, Biswas SB.; Conformational dynamics of DnaB helicase upon DNA and nucleotide binding: analysis by intrinsic tryptophan fluorescence quenching. Biochemistry. 2003 Feb 25;42(7):1910-21.
---...---