As reported by Oshima et al. in 2018, inserting bridged nucleic acids (BNAs) into oligonucleotide probes increase the melting temperature (Tm) of the probes leading to an increase of the delta Tm (Δ Tm) between m6A-containing RNA and unmethylated RNA as compared with DNA probes. The insertion of artificial nucleic acids such as BNAs into oligonucleotides increases both binding affinities to the target nucleic acid and sensitivity for detecting a single base mismatch.
A technique called northeastern blotting allows measuring m6A modifications in RNA. But this method is less efficient, however recent technological developments allow for a more accurate and comprehensive analysis of these modifications in RNA.
Oshima et al. found that the insertion of bridged nucleic acids (BNAs) into DNA probes increases the difference in melting temperature between N6-methyladenosine (m6A)-containing RNA and unmethylated RNA. This approach allowed the researcher to quantify methylation efficiency at m6A sites in E. coli 23S rRNA with high accuracy. The design of the fluorescent BNA probes is illustrated in figure 1.
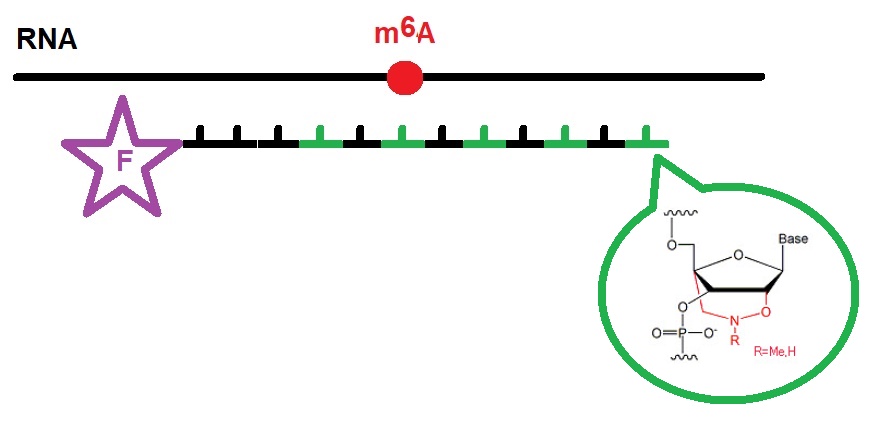
Figure 1: Fluorescent BNA probes for m6A RNAs as designed by Oshima et al. (2018).
N6-Methyladenosine (m6A) modifications are relatively abundant modifications in RNA molecules in ribosomal RNA (rRNA), small nuclear RNA (snRNA) and messenger RNA (mRNA) of various species. N6-Methyladenosine (m6A) modifications of RNA occur in eukaryotes in mRNA, rRNA, tRNA, and microRNA and appear to be reversible since demethylation enzymes are known to be present in various species. RNA modifications affect RNA splicing, translation, degradation, and localization. Modified RNAs have significant roles in transcriptional control, but their role in the regulation of diverse physiological pathways is still unclear. The structure of a RNA duplex containing m6A is illustrated in figure 2.
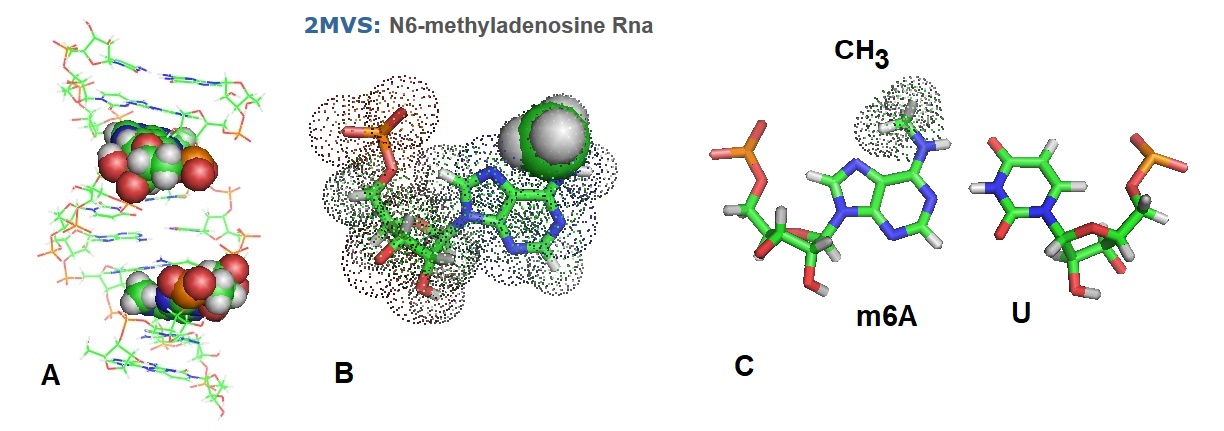
Figure 2: Structure of methylated RNA duplex as reported by Roost et al. in 2015. m6A is oriented anti in a paired duplex, with the methyl group in the anti conformation as well. (A) Structure of the 10 bp duplex with methylated adenosines. (B) Structural model of N6-Methyladenosine as found in RNA. (C) Base pairs alone of m6A/U as found in the duplex.
m6A modifications influence many physiological processes such as circadian rhythms, stem cell pluripotency, fibrosis, triglyceride metabolism, and obesity. m6A modifications are thought to control RNA degradation, localization, and splicing. However, their exact role in pathophysiological states is still unclear. Therefore, one can reason that the investigation of m6A modifications are essential to provide insights into molecular physiology and molecular pathways influenced by this modification.
.
Reference
Roost C, Lynch SR, Batista PJ, Qu K, Chang HY, Kool ET. Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J Am Chem Soc. 2015;137(5):2107-15.
Oshima T , Ishiguro K , Suzuki T , Kawahara Y .; Quantification of methylation efficiency at a specific N6-methyladenosine position in rRNA by using BNA probes. Chem Commun (Camb). 2018 Aug 23;54(69):9627-9630. doi: 10.1039/c8cc03713b.
Wang CY, Lin MH, Su HT. A Method for Measuring RNA N 6-methyladenosine Modifications in Cells and Tissues. J Vis Exp. 2016;(118):54672. Published 2016 Dec 5. doi:10.3791/54672